REST2 vs. Standard MD: A Guide to Enhanced Conformational Sampling for Biomolecular Simulations
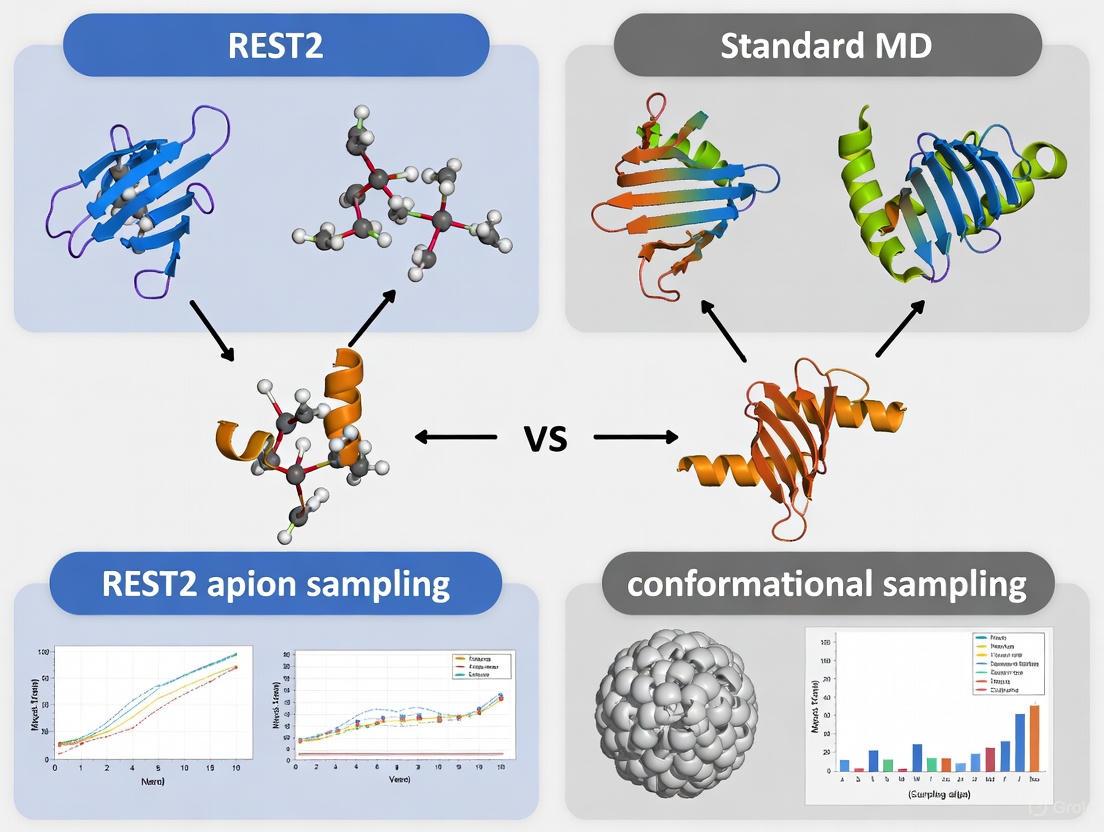
This article provides a comprehensive comparison between Replica Exchange with Solute Tempering 2 (REST2) and standard Molecular Dynamics (MD) for conformational sampling in biomolecular simulations.
REST2 vs. Standard MD: A Guide to Enhanced Conformational Sampling for Biomolecular Simulations
Abstract
This article provides a comprehensive comparison between Replica Exchange with Solute Tempering 2 (REST2) and standard Molecular Dynamics (MD) for conformational sampling in biomolecular simulations. Targeted at researchers and drug development professionals, we explore the foundational principles of enhanced sampling, detailing REST2's Hamiltonian scaling methodology that overcomes the computational limitations of temperature-based replica exchange. The article delivers practical insights into implementing REST2 in modern software like NAMD, examines its application in studying protein folding and intrinsically disordered proteins (IDPs), and addresses critical troubleshooting aspects such as mitigating artificial compaction. Finally, we present a rigorous validation of REST2's performance against standard MD and other sampling methods, evaluating its efficiency in converging thermodynamic averages and exploring conformational landscapes, positioning it as a transformative tool for accelerating drug discovery.
The Sampling Bottleneck: Why Standard MD Struggles with Complex Biomolecular Landscapes
Molecular Dynamics (MD) simulation is a powerful computational tool that provides atomic-level insights into biomolecular processes, from protein folding to drug binding [1] [2]. However, a fundamental limitation plagues conventional MD: the quasi-ergodicity problem. This phenomenon occurs when simulations become trapped in local energy minima—metastable states separated by high free-energy barriers—failing to sample the complete conformational ensemble within accessible simulation timescales [2]. For biologically relevant events occurring on microsecond to millisecond timescales or longer, standard MD with femtosecond integration steps would require >10¹² steps, making comprehensive sampling computationally prohibitive without specialized hardware or advanced algorithms [2].
The consequences of this sampling failure are profound. Without adequate sampling, simulations cannot determine the underlying free energy landscape or correctly estimate the relative populations of different conformational states [2]. This limitation is particularly acute for studying rare events such as protein folding, conformational transitions in allosteric proteins, and ligand unbinding—processes crucial for understanding biological function and designing therapeutics [3] [4]. This article objectively compares the conformational sampling performance between standard MD and the enhanced sampling method Replica Exchange with Solute Tempering (REST2), examining their ability to overcome the quasi-ergodicity problem through quantitative metrics and experimental evidence.
Theoretical Framework: Energy Landscapes and Sampling Barriers
The Rugged Energy Landscape of Biomolecules
Proteins exist not as single rigid structures but as dynamic ensembles of conformations distributed across a high-dimensional free energy landscape according to their Boltzmann-weighted probabilities [2]. This landscape is typically rugged and multifunneled, comprising numerous local minima (metastable states) separated by varying energy barriers [2] [5]. The height of these barriers determines the transition rates between states, with higher barriers leading to exponentially slower transitions in standard MD simulations [2].
For complex biomolecules like intrinsically disordered proteins (IDPs) and metamorphic proteins, the landscape becomes particularly challenging to characterize. IDPs lack a stable folded structure and sample a broad conformational space, while metamorphic proteins adopt multiple distinct folded structures with different functions [5]. Standard MD simulations typically sample only local minima within these complex landscapes, providing an incomplete picture of the conformational ensemble [5].
Molecular Determinants of Trapping
The physical origins of the quasi-ergodicity problem stem from specific molecular interactions that create high energy barriers:
- Hydrophobic collapse and core packing: Burial of hydrophobic residues requires coordinated rearrangement of multiple side chains [2]
- Salt bridge and hydrogen bond networks: Polar interactions often form cooperative networks that must break simultaneously for transitions to occur [2]
- Backbone torsional barriers: Rotations around phi and psi angles encounter significant energy barriers [2]
- Ligand binding "energy cages": In some protein-ligand complexes, conformational rearrangements after initial binding create steric hindrance that traps ligands, requiring substantial energy to dissociate [4]
These molecular features create a rugged landscape where the system spends most of its time vibrating within local minima, rarely sampling transition pathways to other regions of conformational space.
Standard MD vs REST2: Methodological Comparison
Standard Molecular Dynamics Protocol
In standard MD, the system evolves according to Newton's equations of motion in the NVT (canonical) or NPT (isothermal-isobaric) ensemble [6]. The basic algorithm involves:
- Initialization: Input of initial coordinates, velocities, and force field parameters [6]
- Force calculation: Computation of forces on all atoms from bonded and non-bonded interactions [6]
- Integration: Numerical solution of Newton's equations to update positions and velocities [6]
- Output: Periodic saving of trajectories, energies, and other properties [6]
The simulation temperature is maintained using thermostats such as Nosé-Hoover or Berendsen, which rescale velocities to maintain the target temperature [6]. While theoretically sound, this approach suffers from extremely slow barrier crossing in rugged energy landscapes, as the system must wait for rare thermal fluctuations to overcome energy barriers.
REST2 Enhanced Sampling Methodology
Replica Exchange with Solute Tempering (REST2) belongs to the class of generalized ensemble methods that enhance sampling by simulating multiple replicas under different conditions [1] [3]. Unlike standard temperature replica exchange which heats the entire system, REST2 employs a Hamiltonian scaling approach that selectively enhances fluctuations in the solute degrees of freedom while maintaining the solvent at the target temperature [1] [5].
The REST2 protocol involves:
- System partitioning: Division into solute (protein/ligand) and solvent regions [5]
- Hamiltonian scaling: Application of a scaling factor (λ) to the solute potential energy terms across replicas, with λ ranging from 1 (unscaled) to values <1 (effectively "hotter" solute) [5]
- Parallel simulation: Running simultaneous MD simulations for each replica with different scaling factors [3]
- Configuration exchange: Periodic attempts to swap configurations between adjacent replicas based on a Metropolis criterion that preserves detailed balance [3]
The exchange probability between replicas i and j is given by:
$$P{exchange} = min(1, exp[-(βi - βj)(Vi(q^j) - V_j(q^i))])$$
Where β represents the inverse temperature and V the potential energy [3]. This approach allows the solute to effectively sample higher temperatures while maintaining realistic solvent behavior, significantly improving conformational sampling with fewer replicas than temperature-based replica exchange [3] [5].
Table 1: Key Differences Between Standard MD and REST2 Sampling Approaches
| Parameter | Standard MD | REST2 |
|---|---|---|
| Sampling ensemble | Canonical (NVT/NPT) | Generalized ensemble |
| Temperature treatment | Single temperature for entire system | Scaled Hamiltonian for solute regions |
| Replica communication | None (single trajectory) | Multiple replicas with configuration exchange |
| Barrier crossing mechanism | Rare thermal fluctuations | Hamiltonian scaling promotes barrier crossing |
| Computational resource | Single simulation | Multiple parallel simulations with exchange overhead |
| System size limitation | Limited by single simulation cost | Limited by replica number and exchange efficiency |
Quantitative Performance Comparison
Sampling Efficiency Metrics
Multiple studies have quantitatively compared the sampling efficiency of REST2 against standard MD using both model systems and biologically relevant proteins. Efficiency is typically measured by:
- Time to folding: Simulation time required to reach native structure from unfolded states [5]
- Replica mixing rates: Acceptance probabilities for configuration exchanges between replicas [5]
- Free energy barriers: Estimated heights of barriers between conformational states [5]
- Ergodicity measures: Convergence of independent simulations and coverage of conformational space [5]
In benchmark studies on fast-folding proteins like TRP-cage and β-hairpin, REST2 demonstrated significantly improved sampling efficiency compared to standard MD [5].
Comparative Performance Data
Table 2: Quantitative Comparison of Standard MD and REST2 Performance on Model Systems
| Protein System | Standard MD Performance | REST2 Performance | Key Metric |
|---|---|---|---|
| TRP-cage | Folding in ~300 ns (1-2 replicas) | Folding in <100 ns (6/12 replicas) | Time to native structure [5] |
| β-hairpin | Folding in ~300 ns | Folding in <100 ns | Time to native structure [5] |
| Alanine dipeptide | Slow dihedral transitions | Rapid dihedral space coverage | Dihedral transition rates [5] |
| Free energy barrier | ~6 kcal/mol (estimated) | ~2 kcal/mol (matches experimental ~2.1 kcal/mol) | Barrier height estimation [5] |
| Replica mixing | N/A | Efficient solute state exchange | Replica exchange acceptance [5] |
For intrinsically disordered proteins like Histatin-5 and metamorphic proteins like RFA-H, REST2 and its variants provide significantly better agreement with experimental NMR and SAXS data compared to standard MD, without requiring trajectory reweighting [5]. This demonstrates REST2's ability to sample the broad conformational ensembles characteristic of these challenging systems.
Advanced REST2 Applications and Hybrid Methods
Combining REST2 with Generative Models
Recent advances have integrated REST2 with denoising diffusion probabilistic models to further enhance free energy landscape mapping [3]. This hybrid approach treats potential energy as a fluctuating variable within the REST2 framework, then uses diffusion models to learn the joint probability distribution in configuration and rescaled potential energy space [3]. Benchmarking on the mini-protein CLN025 demonstrated that this DDPM-refined REST2 achieves accuracy comparable to temperature replica exchange while requiring fewer replicas [3].
For systems with particularly high barriers, an iterative scheme combining REST2, diffusion models, and importance sampling along known collective variables has been developed to improve resolution in high-barrier regions [3]. Application to the enzyme PTP1B successfully revealed a loop transition pathway consistent with prior biased simulations, showcasing the method's ability to uncover complex transitions with minimal computational overhead compared to conventional replica exchange [3].
Hybrid Tempering Approaches
Further improvements to REST2 led to the development of Replica Exchange with Hybrid Tempering, which differentially and optimally heats both solute and solvent components [5]. Unlike standard REST2, REHT includes additional temperature bias in replicas along with Hamiltonian scaling of the protein solute [5]. This approach accelerates the rewiring of hydration shells that work cooperatively with protein conformational changes, particularly helping overcome entropic barriers [5].
The exchange criteria for REHT incorporates terms for protein-protein, protein-water, and water-water interactions:
$${\Delta}{{{nm}}}\left( {{\rm{REHT}}} \right) = - \left[ \begin{array}{l}({\it{\beta }}{{n}}\lambda {{n}} - {{\beta }}{{m}}\lambda {\rm{m}})\left[ {{{H}}{{{pp}}}\left( {{{X}}{{n}}} \right) - {{H}}{{{pp}}}\left( {{{X}}{{m}}} \right)} \right]\ + \left( {{\it{\beta }}{{n}}\sqrt {\lambda {\rm{n}}} - {\it{\beta }}{{m}}\sqrt {\lambda {\rm{m}}} } \right)\left[ {{{H}}{{{pw}}}\left( {{{X}}{{n}}} \right) - {{H}}{{{pw}}}\left( {{{X}}{{m}}} \right)} \right]\ + \left( {{\it{\beta }}{{n}} - {\it{\beta }}{{m}}} \right)\left[ {{{H}}{{{ww}}}\left( {{{X}}{{n}}} \right) - {{H}}{{{ww}}}\left( {{{X}}_{{m}}} \right)} \right]\end{array} \right]$$
Where Hₚₚ, Hₚ𝓌, and H𝓌𝓌 represent protein-protein, protein-water, and water-water interaction energies, respectively [5]. This hybrid approach has demonstrated significantly improved sampling efficiency across diverse protein types, from simple model systems to complex disordered and metamorphic proteins [5].
Practical Implementation and Research Toolkit
Essential Research Reagents and Software
Table 3: Research Reagent Solutions for REST2 Implementation
| Tool/Category | Specific Examples | Function/Role |
|---|---|---|
| MD Software | GROMACS [6] [1], AMBER [1] [2], CHARMM [1], NAMD [1], GENESIS [1] | Core simulation engines with enhanced sampling capabilities |
| Enhanced Sampling Modules | PLUMED [5] | Plugin for implementing advanced sampling algorithms |
| Force Fields | AMBER [2] [7], CHARMM [2], OPLS [2], GROMOS [2] | Molecular mechanical parameter sets for biomolecules |
| Specialized Force Fields | RNA-specific χOL3 [7] | Domain-specific parameters for accurate RNA simulation |
| Analysis Tools | MDTraj, MDAnalysis, VMD | Trajectory analysis and visualization |
| Hybrid Methods | Denoising Diffusion Probabilistic Models (DDPMs) [3] | Generative models for refining free energy landscapes |
Experimental Workflow and Best Practices
Diagram 1: REST2 simulation workflow showing the parallel replica approach with periodic configuration exchanges.
Successful implementation of REST2 requires careful attention to several practical considerations:
- Replica number and spacing: The number of replicas should be sufficient to ensure reasonable exchange probabilities (>20% typically recommended) between adjacent replicas [5]
- Hamiltonian scaling range: λ values typically range from 1.0 (unscaled) down to ~0.6, adjusted based on system size and complexity [5]
- Exchange frequency: Configuration exchange attempts typically occur every 1-10 ps, balancing communication overhead with conformational decorrelation [5]
- Convergence assessment: Ergodicity should be verified by comparing conformational distributions from different trajectory segments and monitoring replica diffusion through parameter space [5]
For RNA systems, recent CASP15 benchmarking suggests MD refinement works best for stabilizing already high-quality models rather than correcting poor initial structures, with optimal simulation lengths typically 10-50 ns [7].
The quasi-ergodicity problem presents a fundamental challenge in biomolecular simulation that standard MD cannot adequately address for many biologically relevant processes. REST2 and its variants provide a robust solution by selectively enhancing solute fluctuations while maintaining realistic solvent behavior, enabling more efficient exploration of complex energy landscapes.
Quantitative comparisons demonstrate REST2's superiority in sampling speed, barrier crossing efficiency, and convergence for diverse systems ranging from fast-folding model proteins to complex disordered and metamorphic proteins. The continued development of hybrid approaches combining REST2 with generative models and other enhanced sampling techniques promises further advances in mapping biomolecular free energy landscapes with unprecedented resolution and efficiency.
For researchers studying conformational dynamics, binding mechanisms, or allosteric regulation, REST2 offers a compelling alternative to standard MD when facing the quasi-ergodicity problem. Its ability to sample functionally relevant states separated by significant energy barriers makes it particularly valuable for drug discovery applications where understanding rare transitions can illuminate mechanisms of action and opportunities for therapeutic intervention.
Molecular dynamics (MD) simulation serves as a computational microscope, enabling researchers to study biomolecular motions at atomic resolution. However, the potential energy landscape of biomolecules is characterized by numerous energy minima and high barriers, making adequate conformational sampling a significant challenge. Enhanced sampling techniques are therefore essential for studying processes like protein folding, ligand binding, and conformational changes in intrinsically disordered proteins (IDPs). Among these techniques, Temperature Replica Exchange MD (T-REMD) has been widely adopted, but its application to large, solvated systems is severely limited by a fundamental scaling problem. This guide objectively compares the performance of T-REMD with its more efficient alternative, Replica Exchange with Solute Tempering (REST2), focusing specifically on their scalability with system size and their efficacy in conformational sampling for drug development research.
How Replica Exchange Methods Work: Fundamental Principles
Replica exchange molecular dynamics is a generalized ensemble method designed to overcome energy barriers and escape local minima, which are common obstacles in conventional MD simulations. The core principle involves running multiple simultaneous copies (replicas) of the system under different thermodynamic conditions.
T-REMD and REST2 Workflows
Diagram 1: Comparative workflows of T-REMD and REST2. T-REMD scales the entire system's temperature, while REST2 uses Hamiltonian scaling to target only the solute region, dramatically improving efficiency for large solvated systems.
In T-REMD, replicas are run at different temperatures, and periodic exchange attempts between adjacent temperatures are made based on the Metropolis criterion [8]. This enables random walks in temperature space, helping the system overcome energy barriers. In contrast, REST2 applies Hamiltonian rescaling to achieve effective tempering only in selected solute regions while the solvent remains at a constant temperature for all replicas [9]. This fundamental difference in approach has profound implications for computational efficiency and practical applicability.
The System Size Challenge: Quantitative Analysis of T-REMD Limitations
Mathematical Framework of Poor Scaling
The primary limitation of T-REMD is its poor scaling with system size. The number of replicas required to maintain adequate exchange probabilities grows with the square root of the number of degrees of freedom in the system. For a biomolecular system with N atoms, the number of replicas needed to cover a given temperature range scales as O(√N) [9]. This relationship becomes prohibitively expensive for large systems, particularly those with explicit solvent representation.
Table 1: Replica Requirements for T-REMD vs. REST2
| System Description | Total Atoms | T-REMD Replicas Required | REST2 Replicas Required | Computational Savings |
|---|---|---|---|---|
| p53 N-terminal domain (IDP) | ~72,000 | >100 [9] | 16 [9] | ~84% reduction |
| Small globular protein | ~30,000 | ~50 [8] | 12-16 [9] | ~70% reduction |
| Peptide-water system | ~10,000 | ~24 [8] | 8-12 [9] | ~60% reduction |
This mathematical relationship has severe practical consequences. For the disordered N-terminal domain of p53 (p53-NTD, residues 1-61) solvated in approximately 72,000 atoms, a T-REMD simulation would require over 100 replicas to achieve acceptable exchange rates (~20%) between 298 K and 500 K [9]. In contrast, the same system simulated with REST2 requires only 16 replicas to cover the same temperature range while maintaining approximately 25% acceptance rates [9].
Physical Origin of the Scaling Problem
The poor scaling of T-REMD originates from the statistical mechanical relationship between system size and energy fluctuations. The probability of exchanging two replicas at temperatures Ti and Tj depends on their potential energy distributions, with the overlap between these distributions determining the acceptance rate. As system size increases, the energy distributions become narrower relative to their means, reducing the overlap between adjacent replicas and consequently lowering exchange probabilities [1] [9].
In biomolecular simulations with explicit solvent, the total energy is dominated by solvent-solvent interactions rather than solute-solute or solute-solvent interactions. Conventional T-REMD wastes computational resources by heating the entire system, including the solvent, when often only the conformational sampling of the solute is of interest [1].
REST2: A Scalable Alternative for Biomolecular Simulations
Fundamental Mechanism and Theoretical Basis
Replica Exchange with Solute Tempering (REST2) addresses the scaling problem by targeting the enhanced sampling specifically to regions of interest. The method employs Hamiltonian rescaling to create an effective temperature ladder for selected solute regions while maintaining the solvent at a constant temperature across all replicas [9].
The scaled Hamiltonian in REST2 is defined as:
[ Em^{REST2} = \lambdam^{pp}E{pp} + \lambdam^{pw}E{pw} + \lambda^{ww}E{ww} ]
Where (E{pp}), (E{pw}), and (E{ww}) represent solute-solute, solute-solvent, and solvent-solvent interaction energies, respectively. The scaling factors are set as (\lambdam^{pp} = \betam/\beta0), (\lambdam^{pw} = \sqrt{\betam/\beta0}), and (\lambda^{ww} = 1), with (\betam = 1/kBTm) and (\beta0 = 1/kBT_0) [9].
This formulation means only interactions related to the solute contribute to the Metropolis criteria for replica exchange, dramatically reducing the number of degrees of freedom involved in exchange attempts and consequently requiring fewer replicas to cover the same effective temperature range.
Experimental Validation and Performance Metrics
Table 2: Sampling Efficiency Comparison for IDP Systems
| Performance Metric | T-REMD | REST2 | Experimental Reference |
|---|---|---|---|
| Replica count for p53-NTD | >100 | 16 | [9] |
| Acceptance rate | ~20% (estimated) | ~25% | [9] |
| Conformational convergence | Limited without extensive sampling | Improved with equivalent computational resources | [10] |
| Sampling of rare events | Possible but computationally expensive | Enhanced through targeted tempering | [3] |
| Force field validation | Used in implicit and explicit solvent optimizations | Applied to explicit solvent IDP simulations | [9] |
REST2 has demonstrated particular effectiveness for studying intrinsically disordered proteins (IDPs), which sample heterogeneous conformational ensembles and require extensive sampling. HREMD (closely related to REST2) produces configurational ensembles consistent with SAXS, SANS, and NMR experiments for IDPs with varying sequence characteristics, including Histatin 5 (24 residues) and Sic 1 (92 residues) [10]. The agreement with multiple experimental techniques without biasing or reweighting the simulations confirms the method's validity for generating accurate structural ensembles [10].
Practical Implementation and Protocol Design
Research Reagent Solutions: Computational Tools
Table 3: Essential Research Resources for Replica Exchange Simulations
| Resource Category | Specific Tools | Function and Application |
|---|---|---|
| MD Simulation Software | GROMACS [1], AMBER [1], NAMD [1], CHARMM [1], GENESIS [1] | Core simulation engines implementing T-REMD and REST2 algorithms |
| Enhanced Sampling Methods | REMD [8], REST2 [9], HREMD [10], gREST [1], ALSD [1] | Specialized algorithms for improved conformational sampling |
| Force Fields | Amber ff03ws [10], Amber ff99SB-disp [10], CHARMM36 [11] | Energy functions parameterized for proteins and nucleic acids |
| Analysis Tools | PyEMMA [8], MSMBuilder [8], SHIFTX2 [10] | Processing trajectories and calculating experimental observables |
| Validation Methods | SAXS/SANS [10], NMR chemical shifts [10] | Experimental techniques for validating computational ensembles |
Detailed Experimental Protocol for REST2
Implementing REST2 requires careful attention to several technical aspects:
System Preparation: The biomolecular system must be solvated in an appropriate water box with sufficient padding to accommodate conformational fluctuations. For IDPs, this is particularly important as they can sample extended conformations [10].
Replica Parameterization: The number of replicas and their effective temperature spacing should be optimized for the specific system. For a typical protein system, 12-24 replicas are sufficient with REST2, compared to 50-100+ with T-REMD [9].
Hamiltonian Scaling: The scaling factors for the solute-solute ((\lambdam^{pp})) and solute-solvent ((\lambdam^{pw})) interactions must be set according to the REST2 protocol: (\lambdam^{pp} = \betam/\beta0) and (\lambdam^{pw} = \sqrt{\betam/\beta0}) [9].
Simulation Parameters: Exchange attempts should occur every 1-2 ps, with simulation lengths of 500 ns per replica or longer for larger systems, as used in successful HREMD studies of IDPs [10].
Validation: The resulting ensembles should be validated against experimental data such as SAXS curves and NMR chemical shifts to ensure physical relevance [10].
Limitations and Recent Advancements
Known Limitations of REST2
Despite its advantages, REST2 has limitations. The method can promote artificial protein conformational collapse at high effective temperatures, particularly for larger IDPs [9]. This collapse can lead to replica segregation in the effective temperature space, hindering sampling of large-scale conformational changes [9]. Additionally, the scaling of solute-solvent interactions in REST2 intentionally weakens these interactions at higher temperatures, which was designed to promote refolding of small proteins but may not be optimal for studying extended conformational ensembles of IDPs [9].
Emerging Alternatives and Improvements
Recent research has addressed these limitations through method refinements:
REST3: A new protocol that recalibrates the scaling of solute-solvent van der Waals interactions to reproduce appropriate levels of protein chain expansion at high effective temperatures, eliminating exchange bottlenecks and improving temperature random walk [9].
Hybrid Approaches: Combining REST2 with diffusion-based generative models (DDPM) enhances the mapping of conformational free-energy landscapes and improves sampling of high-barrier regions [3].
Generalized REST (gREST): Extends the approach to allow selective enhancement of arbitrary regions within the solute, not just the entire biomolecule [1].
Diagram 2: Evolution of replica exchange methods. The methodological development path from identifying the T-REMD scaling problem through REST2 development to its recent refinements and future directions incorporating machine learning approaches.
The system size challenge fundamentally limits the application of traditional T-REMD to biologically relevant systems with explicit solvent. REST2 and its variants address this limitation through Hamiltonian rescaling that targets enhanced sampling to regions of interest, reducing replica requirements by 60-84% compared to T-REMD. While REST2 has proven particularly valuable for studying intrinsically disordered proteins and large biomolecular systems, researchers should be aware of its tendency to promote artificial compaction in some systems and consider recent improvements like REST3 or hybrid approaches combining REST2 with machine learning for challenging sampling problems. As biomolecular simulations continue to tackle increasingly complex systems, the development and refinement of targeted enhanced sampling methods like REST2 will remain crucial for bridging computational and experimental studies in structural biology and drug discovery.
Molecular dynamics (MD) simulations are powerful tools for studying the movement and interactions of biological molecules, such as proteins, at an atomic level. A significant challenge, however, is that these molecules often undergo functional conformational changes on timescales that are computationally expensive—sometimes impossible—to simulate with standard MD. Enhanced sampling methods were developed to overcome this hurdle by accelerating the exploration of a molecule's energy landscape. Among these, Replica Exchange with Solute Tempering (REST2) stands out as an efficient and widely adopted method. REST2 belongs to a class of enhanced sampling techniques known as Hamiltonian Replica Exchange, which modifies the energy function of the system to improve sampling efficiency. This guide provides a objective comparison of REST2 against other prominent enhanced sampling methods, supported by recent experimental data and implementation protocols.
What is REST2? Core Principles and Mechanism
Replica Exchange with Solute Tempering 2 (REST2) is an enhanced sampling algorithm designed to efficiently explore the conformational space of a biomolecule, such as a protein or a peptide. Its core innovation is to focus the sampling acceleration on a "solute" region of interest—for example, a protein—while treating the surrounding solvent environment more efficiently.
The method operates on the following key principles [12] [13] [3]:
- Hamiltonian Replica Exchange: Unlike Temperature Replica Exchange (TREM), which runs parallel simulations at different physical temperatures, REST2 runs all replicas at the same physical temperature (typically the ambient temperature of interest). Instead, the energy function (Hamiltonian) is scaled differently in each replica.
- Selective Scaling: The scaling of the Hamiltonian is applied selectively to the interactions within the solute (e.g., protein-protein interactions) and between the solute and the solvent. The solvent-solvent interactions remain unscaled. This creates a ladder of replicas where the solute effectively experiences different "temperatures," encouraging it to escape local energy minima.
- Replica Exchange: Periodic swaps between adjacent replicas are attempted based on a Metropolis criterion. This allows a conformation that has been "heated up" and unfolded in one replica to be transferred to a colder replica, where it can refold, thus efficiently traversing the energy landscape.
The diagram below illustrates the logical workflow and key concepts of the REST2 method.
Performance Comparison: REST2 vs. Alternative Methods
REST2 is one of several strategies to enhance sampling. The table below provides a high-level comparison of its approach against other common methods.
Table 1: Comparison of Enhanced Sampling Methodologies
| Method | Core Principle | Key Advantage | Key Limitation |
|---|---|---|---|
| REST2 | Hamiltonian replica exchange with scaling applied to solute-solute and solute-solvent interactions. | More efficient than TREM for large systems; solvent remains at ambient temperature. | Requires communication between parallel replicas; performance can be hindered on heterogeneous computing clusters [12]. |
| Temperature REMD (TREM) | Parallel simulations at different temperatures with exchanges. | Conceptually simple; effective for small systems. | Number of replicas grows with system size, becoming computationally prohibitive for large biomolecules [12]. |
| Simulated Tempering (ST) | A single simulation that updates its temperature based on a Metropolis criterion. | No communication between parallel runs required; efficient on heterogeneous hardware [12]. | Can be less efficient than REST2 for biomolecular systems, requiring more temperature "rungs" [12]. |
| Simulated Solute Tempering 2 (SST2) | A combination of ST and REST2; a single simulation updates its scaled Hamiltonian. | Achieves comparable or superior sampling to REST2 with fewer replicas; no inter-replica communication [12]. | A newer method, less established in community-wide usage compared to REST2. |
| Biased Sampling (e.g., Metadynamics) | Applies a bias potential along pre-defined Collective Variables (CVs) to push the system out of energy minima. | Can be extremely efficient if a good CV (e.g., a true reaction coordinate) is known [14]. | Performance is entirely dependent on the correct choice of CVs, which is often non-trivial [14]. |
| Generative AI Models (e.g., DDPM) | Machine learning models trained on simulation data to generate new, statistically likely conformations. | Can generate novel conformations and enhance sampling with significant computational savings [15] [3]. | Limited by the quality of training data; cannot discover states not already partially sampled in the input simulations [15] [3]. |
Quantitative Performance Data
Theoretical comparisons are best validated with experimental data. A recent study benchmarked REST2 against ST, SST1, and SST2 on two small model proteins: chignolin CLN025 and Trp-Cage [12]. The simulations were run starting from both folded (F) and unfolded (U) states to assess sampling efficiency and the ability to recover correct folding thermodynamics.
Table 2: Experimental Sampling Efficiency on Model Systems [12]
| System | Sampling Method | Simulation Length per Replica | Number of Replicas | Key Finding |
|---|---|---|---|---|
| Chignolin CLN025 | ST | 10 μs | 20 | Serves as a baseline but requires a high number of replicas. |
| SST1 | 10 μs | 10 | Improved over ST but may be limited for large conformational changes. | |
| REST2 | 1 μs | 10 | Achieved efficient sampling with shorter simulation times. | |
| SST2 | 10 μs | 10 | Achieved comparable or superior sampling to REST2 in this test. | |
| Trp-Cage | ST | 40 μs | 20 | Requires long simulation times and many replicas. |
| REST2 | Data not specified | 10 | Demonstrated high efficiency for sampling folded states. | |
| SST2 | 40 μs | 10 | Performance comparable to REST2. |
This data demonstrates that REST2 can achieve high sampling efficiency with fewer replicas than traditional ST and potentially shorter simulation times than other methods, making it a robust and practical choice.
Experimental Protocols and Workflows
To ensure reproducibility and provide a clear guide for researchers, this section outlines a general protocol for setting up and running a REST2 simulation, based on its standard implementation.
A Generic REST2 Simulation Workflow
The following diagram details the key steps involved in a typical REST2 simulation, from system preparation to analysis.
Detailed Methodology for a REST2 Study
The following protocol is synthesized from studies that have successfully employed REST2, such as the investigation of the disordered protein ELF3 [13].
System Preparation:
- Structure Source: Obtain the initial atomic coordinates of the biomolecule. This can come from experimental structures (Protein Data Bank, PDB) or from computational models (e.g., AlphaFold2 prediction) [13].
- Solvation: Use software like GROMACS, NAMD, or OpenMM to place the solute in a simulation box filled with explicit water molecules (e.g., TIP3P model).
- Neutralization: Add ions (e.g., Na⁺, Cl⁻) to neutralize the system's net charge and, optionally, to achieve a physiological ion concentration.
Parameter Setting:
- Solute/Solvent Definition: Clearly define which atoms belong to the solute (the region of interest) and which to the solvent (water and ions).
- Replica Ladder: Choose the number of replicas (e.g., 10) and the scaling factors (λ) for the Hamiltonian. These factors typically range from 1 (unscaled, reference system) to a value that effectively corresponds to a high temperature for the solute. The values are chosen to ensure a sufficient exchange probability (e.g., 20-30%) between adjacent replicas.
Simulation Execution:
- Equilibration: Run a standard equilibration protocol for the unscaled system, including energy minimization, heating to the target temperature (e.g., 300 K), and equilibration of density.
- Production REST2: Launch the parallel REST2 simulation. Each replica runs with its own scaled Hamiltonian. After a fixed number of steps (e.g., every 1-2 ps), the coordinates and energies of adjacent replicas are compared, and a swap is attempted based on the Metropolis Monte Carlo criterion.
Data Analysis:
- Trajectory Analysis: Analyze the combined trajectories from all replicas, focusing on the replica at the reference temperature (λ=1) or using reweighting techniques to compute properties at the target temperature.
- Free Energy Calculation: Use methods like the Multistate Bennett Acceptance Ratio (MBAR) to compute conformational free energies and reconstruct free energy surfaces from the REST2 simulation data [3].
Successful implementation of REST2 requires a combination of software, force fields, and computational resources. The following table lists key "research reagents" for this field.
Table 3: Essential Tools and Resources for REST2 Simulations
| Item | Function in Research | Example Software / Database |
|---|---|---|
| MD Simulation Engine | Software that performs the numerical integration of Newton's equations of motion and implements the REST2 algorithm. | GROMACS [12], NAMD [12], OpenMM, AMBER |
| Molecular Viewing Software | Used to visualize initial structures, simulation trajectories, and final conformations. | VMD, PyMol, UCSF Chimera |
| Force Field | A set of empirical parameters that describe the potential energy of the system; critical for accuracy. | CHARMM36 [13], AMBER ff19SB, OPLS-AA |
| Water Model | Represents the behavior of solvent water molecules in the simulation. | TIP3P [13], SPC/E, TIP4P |
| Structure Database | Source for initial experimental structures of proteins and complexes. | Protein Data Bank (PDB) [12] |
| Analysis Tools | Software packages for processing MD trajectories to compute metrics like RMSD, radius of gyration, and free energies. | MDTraj, PyEMMA, MDAnalysis, GROMACS analysis tools |
| High-Performance Computing (HPC) | Computational clusters (CPUs/GPUs) are essential for running the multiple, parallel replicas in a timely manner. | Local clusters, National supercomputing centers, Cloud computing |
Within the landscape of enhanced sampling methods, REST2 has established itself as a powerful and efficient approach for studying biomolecular conformational dynamics. Its key strength lies in its Hamiltonian replica exchange scheme, which focuses computational effort on the solute, allowing for effective exploration of complex energy landscapes with fewer resources than temperature-based replica exchange. As demonstrated by benchmark studies, REST2 achieves performance comparable to or exceeding that of other advanced methods like ST and SST1. While newer techniques, including generative AI and combined approaches, are emerging as promising tools, REST2 remains a well-validated, practical, and highly reliable choice for researchers investigating processes from protein folding and ligand binding to the dynamics of intrinsically disordered proteins. Its implementation in major MD software packages ensures its continued accessibility and utility for the scientific community.
Molecular dynamics (MD) simulations are indispensable for probing biomolecular structure and dynamics, yet their utility is often limited by the problem of quasi-ergodicity—the inability to adequately sample conformational space due to high energy barriers separating local minima. Generalized ensemble methods, particularly the Temperature Replica Exchange Method (TREM), address this by running multiple replicas at different temperatures and permitting configuration exchanges. However, TREM's scalability is poor; the number of required replicas scales with the square root of the system's degrees of freedom (√f), making it prohibitively expensive for large solvated biomolecules where most degrees of freedom belong to the solvent [16].
Replica Exchange with Solute Tempering (REST1) emerged as a transformative solution, drastically reducing the number of necessary replicas by effectively "heating" only the solute while the solvent remains "cold." This innovation meant the number of replicas now scaled with the square root of the solute's degrees of freedom (√fp), offering significant computational savings [16]. Despite this advance, applications to systems with large-scale conformational changes, like trpcage and β-hairpin, revealed limitations in sampling efficiency, with replicas often becoming trapped in folded or extended states [16].
This guide examines the critical evolution from REST1 to its successor, REST2 (Replica Exchange with Solute Scaling). We will objectively compare their performance against each other and standard TREM, supported by experimental data and detailed methodologies, to provide researchers and drug development professionals with a clear understanding of their capabilities within conformational sampling research.
Core Mechanism: A Fundamental Shift in Hamiltonian Scaling
The fundamental difference between REST1 and REST2 lies in their treatment of temperatures and potential energy surfaces across replicas. This shift in strategy is the source of REST2's enhanced performance.
The REST1 Framework
In REST1, different replicas run at different physical temperatures (Tm). The potential energy function for a replica at temperature Tm is deformed as follows [16]: EmREST1(X) = Epp(X) + ((β0 + βm) / 2βm) Epw(X) + (β0 / βm) Eww(X)
Here, X represents the system configuration, βm = 1/kBTm, and T0 is the target temperature. The energy is decomposed into solute-solute (Epp), solute-solvent (Epw), and solvent-solvent (Eww) components. While Eww disappears from the replica exchange acceptance probability, the protein intramolecular potential (Epp) remains unscaled. Consequently, replicas still navigate the full, unmodified energy landscape of the solute, complete with its high barriers [16].
The REST2 Innovation
REST2 represents a paradigm shift. All replicas are run at the same physical temperature, T0, but each replica experiences a differently scaled potential energy surface [16]: EmREST2(X) = (βm / β0) Epp(X) + √(βm / β0) Epw(X) + Eww(X)
A critical change is the scaling of the solute intramolecular energy, Epp, by a factor (βm / β0) that is less than 1 for replicas with Tm > T0. This scaling directly reduces the energy barriers between different solute conformations, making transitions more frequent. Furthermore, the scaling factor for the solute-solvent interaction energy, Epw, is changed from (β0 + βm)/2βm in REST1 to √(βm / β0) in REST2. This minor-seeming change, coupled with the scaling of Epp, enables a more efficient random walk in conformational space [16].
Table 1: Comparison of Hamiltonian Scaling in REST1 and REST2
| Feature | REST1 (Replica Exchange with Solute Tempering) | REST2 (Replica Exchange with Solute Scaling) |
|---|---|---|
| Replica Temperatures | Different physical temperatures (Tm) | Same physical temperature (T0) for all replicas |
| Scaling Strategy | Deformed potential energy at different temperatures | Different potential energy surfaces at one temperature |
| Solute Energy (Epp) | Unscaled: Full barriers remain | Scaled by (βm/β0): Barriers are lowered |
| Solute-Solvent Energy (Epw) | Scaled by (β0 + βm)/2βm | Scaled by √(βm / β0) |
| Solvent Energy (Eww) | Scaled by (β0 / βm) | Unscaled |
| Primary Enhancement | Effective heating of the solute | Direct scaling down of solute energy barriers |
The logical relationship between the different enhanced sampling methods and the key improvements introduced by REST2 is summarized in the diagram below.
Diagram 1: Evolution from MD to REST2 and its key advantages.
Performance Comparison: Experimental Data and Quantitative Benchmarks
The theoretical advantages of REST2 translate into measurable performance gains. Benchmarking studies on small proteins like the trpcage miniprotein and a β-hairpin provide direct, quantitative comparisons of the sampling efficiency between TREM, REST1, and REST2.
Key Performance Metrics and Experimental Protocol
In a foundational study, the folding landscapes of trpcage (a 20-residue protein) and a β-hairpin were simulated using TREM, REST1, and REST2 [16]. The core metrics for comparison were:
- Sampling Efficiency: The rate at which the simulation explores different conformational states, particularly the transitions between folded and unfolded states.
- Replica Exchange Acceptance Probability: The probability that a proposed swap of configurations between two adjacent replicas is accepted. Higher probabilities lead to better random walks across the temperature ladder.
- Computational Cost: Primarily determined by the number of replicas (CPUs) required to achieve adequate exchange probabilities.
The experimental workflow for such a comparative study is outlined below.
Diagram 2: General workflow for comparing TREM, REST1, and REST2.
Comparative Performance Data
The results from the folding studies clearly demonstrate REST2's superiority. The quantitative outcomes are summarized in the table below.
Table 2: Performance Comparison of TREM, REST1, and REST2 on Protein Folding
| Performance Metric | TREM | REST1 | REST2 |
|---|---|---|---|
| Number of Replicas (CPUs) Required | High (Scales with √f) | Reduced (Scales with √fp) | Reduced (Scales with √fp) |
| Replica Exchange Acceptance Probability | Baseline | Lower than REST2 | Significantly Higher |
| Sampling of Folded/Unfolded Transitions | Baseline | Inefficient; prone to trapping | Highly Efficient |
| CPU Time for ab initio Folding | High | Lower than TREM, but inefficient for large changes | Greatly Reduced |
| Key Limitation | Poor system size scaling | Inefficient for large conformational changes | - |
The critical finding was that while both REST1 and REST2 reduce the number of required CPUs compared to TREM, REST2 "greatly increases the sampling efficiency over REST1" [16]. Specifically, for trpcage and the β-hairpin, REST1 simulations showed poor exchange between folded and unfolded states, whereas REST2 facilitated efficient transitions across this conformational divide. The improvement stems from two factors: the direct lowering of intramolecular energy barriers and a more favorable replica exchange acceptance criterion that benefits from an approximate cancellation between Epp and the scaled Epw terms in REST2 [16].
The Scientist's Toolkit: Essential Research Reagents and Methods
To effectively implement and utilize REST2 in conformational sampling research, a specific set of computational tools and methods is essential. The following table details key components of the modern REST2 research toolkit.
Table 3: Research Reagent Solutions for Enhanced Sampling with REST2
| Tool/Method | Category | Primary Function |
|---|---|---|
| REST2 (Hamiltonian Replica Exchange) | Enhanced Sampling Method | Accelerates conformational exploration by scaling solute Hamiltonian terms, reducing the number of replicas needed vs. TREM [16] [1]. |
| Denoising Diffusion Probabilical Models (DDPM) | Generative AI / Analysis | Refines sampling data from REST2 simulations; learns joint probability distributions to generate new configurations and improve free-energy surface resolution [17] [3]. |
| Weighted Ensemble (WE) Sampling | Enhanced Sampling / Benchmarking | Enables efficient exploration of rare events by using progress coordinates (e.g., from TICA) to run parallel, weighted trajectories; useful for benchmarking MD methods [18]. |
| Zero-Multipole Summation Method (ZMM) | Electrostatic Calculation | Provides efficient electrostatic energy calculation under assumption of local neutrality; can be combined with GEPS like REST2 for faster simulations [1]. |
| gREST / ALSD | Generalized Ensemble Method | Allows selective enhancement of conformational sampling in specific regions of a system (e.g., a protein loop or ligand), building on the REST2 concept [1]. |
The evolution from REST1 to REST2 represents a critical, methodology-level advancement in biomolecular simulation. By shifting from a pure temperature-based paradigm to a Hamiltonian scaling one, REST2 directly addresses the dual challenges of system-size scalability and inefficient sampling of large-scale conformational changes. Experimental benchmarks consistently show that REST2 achieves the computational efficiency of REST1 while surpassing its sampling power, delivering performance that is competitive with—and often superior to—standard TREM at a fraction of the cost.
The utility of REST2 continues to grow, forming the foundation for next-generation sampling strategies. Its integration with generative AI models like Denoising Diffusion Probabilistic Models (DDPMs) demonstrates how modern machine learning can leverage the broad exploration provided by REST2 to refine free-energy landscapes and uncover high-barrier transition pathways [17] [3]. Furthermore, the development of generalized ensemble methods for partial systems (GEPS) that allow selective scaling of specific protein regions or energy terms is a direct descendant of the REST2 philosophy [1]. For researchers and drug developers focused on understanding protein folding, enzyme mechanisms, and ligand binding, REST2 remains an indispensable tool in the computational arsenal, enabling more realistic and comprehensive simulations of biological processes.
Implementing REST2: A Practical Framework for Efficient Biomolecular Sampling
Replica Exchange with Solute Tempering 2 (REST2) is an advanced molecular dynamics (MD) sampling algorithm designed to overcome the significant computational limitations of conventional simulation methods. In the study of biomolecular systems, particularly those involving large-scale conformational changes like protein folding or the dynamics of intrinsically disordered proteins (IDPs), achieving sufficient sampling of the energy landscape is a major challenge with standard temperature-based replica exchange (T-REMD), as the number of required replicas scales with the square root of the total number of atoms in the system, making simulations of large solvated systems prohibitively expensive [16] [19]. REST2 addresses this fundamental issue by transforming the Hamiltonian—the mathematical function describing the system's total energy—for each replica rather than simply changing the temperature. This innovative approach allows the enhanced sampling effort to be focused primarily on the solute molecule, while the solvent remains "cold," leading to a drastic reduction in the number of replicas needed and a consequent increase in computational efficiency [20] [16] [19].
The core principle of REST2 lies in its intelligent scaling of different components of the potential energy. The method is founded on the Hamiltonian Replica Exchange (H-REM) framework, where all replicas are simulated at the same physical temperature, but each replica experiences a differently scaled version of the potential energy function [16]. This strategic scaling effectively lowers the energy barriers within the solute, enabling more rapid crossing between different conformational states during the simulation. The resulting performance improvement is substantial; studies have confirmed that REST2 achieves sampling efficiency comparable to other advanced methods like bias-exchange metadynamics (BEMD) and T-REMD, but with far greater computational efficiency and without introducing biases from pre-defined collective variables [20]. This makes REST2 a powerful tool for quantitative biophysical simulations, including peptide folding-unfolding transitions, absolute binding affinity calculations, and free energy landscape exploration [19].
Mathematical Decomposition of the REST2 Hamiltonian
The REST2 algorithm achieves its efficiency through a specific, non-uniform scaling of the potential energy terms. The total potential energy of a molecular system in an explicit solvent can be conceptually partitioned into three primary components:
- Solute-solute interaction energy (E_pp): This term encompasses all intra-molecular interactions within the solute molecule, including bonded interactions (bonds, angles, dihedrals) and non-bonded interactions (van der Waals and electrostatics) between its atoms.
- Solute-solvent interaction energy (E_pw): This term represents the non-bonded interactions between the atoms of the solute and the atoms of the surrounding solvent molecules.
- Solvent-solvent interaction energy (E_ww): This term includes all interactions among the solvent molecules themselves.
In REST2, the potential energy function for a given replica m is defined by applying distinct scaling factors to these components [16] [19]:
Where:
- X represents the configuration (atomic coordinates) of the entire system.
- βm = 1/kB T_m, where
k_Bis Boltzmann's constant andT_mis the effective temperature assigned to the solute for replica m. - β0 = 1/kB T_0, where
T_0is the target physical temperature of the simulation (e.g., 300 K).
The following diagram illustrates the logical relationship between the scaling factors and the resulting effective energy landscape for the solute:
This scaling scheme ensures that the solvent-solvent interactions (E_ww) remain entirely unscaled, preserving the realistic behavior of the solvent at the target temperature T_0. The solute-solute term (E_pp) is scaled by a factor less than 1 for replicas with T_m > T_0, which directly lowers the energy barriers of the solute's internal potential, facilitating transitions between conformational states. The solute-solvent term (E_pw) is scaled by the square root of that factor, a choice that proves critical for maintaining high acceptance probabilities for exchanges between replicas, as it leads to a beneficial partial cancellation of energy fluctuations in the acceptance criterion [16].
Comparative Performance: REST2 vs. Alternative Sampling Methods
To objectively evaluate REST2's performance, it must be compared against other widely used sampling techniques. The key alternatives include standard Temperature Replica Exchange (T-REMD) and the original version of Replica Exchange with Solute Tempering (REST1). The comparison can be based on several critical metrics: computational efficiency, sampling effectiveness, and applicability to different biological problems.
Table 1: Comparative Analysis of REST2 vs. Other Sampling Methods
| Method | Key Principle | Scaling of Replicas with System Size | Computational Efficiency | Sampling Bias | Ideal Use Case |
|---|---|---|---|---|---|
| REST2 | Hamiltonian scaling of solute energy terms [16] | √(f_p) [16] | High (Fewer replicas needed) [19] | No predefined bias [20] | Peptide folding, IDP conformational landscapes, protein-ligand binding [20] [19] |
| T-REMD | Entire system simulated at different temperatures [20] | √(f) [16] | Low (Many replicas needed for large systems) [19] | No predefined bias | Small proteins and peptides in explicit solvent |
| REST1 | Hamiltonian scaling with (β0+βm)/2βm for Epw [16] | √(f_p) [16] | Moderate (Less efficient than REST2 for large changes) [16] | No predefined bias | Systems with modest conformational changes |
| BEMD | History-dependent bias on collective variables [20] | Independent of system size | Variable (Depends on CV choice) | High (Biased by user-defined CVs) [20] | Systems with known, well-defined reaction coordinates |
Legend: f = total degrees of freedom in the system; f_p = degrees of freedom of the solute.
Quantitative benchmarks highlight REST2's advantages. In a study on the intrinsically disordered protein amylin, REST2 yielded results "qualitatively consistent with experiments and in quantitative agreement with other sampling methods, however far more computationally efficiently and without any bias" [20]. Furthermore, comparative folding simulations of the Trp-cage mini-protein and a β-hairpin demonstrated that REST2 "greatly reduces the number of CPUs required by regular replica exchange and greatly increases the sampling efficiency over REST1" [16]. This performance gain is attributed to REST2's more effective lowering of intra-solute energy barriers and its optimized scaling of the solute-solvent interaction, which together enhance the sampling of large-scale conformational transitions.
Experimental Protocols and Validation
The implementation and validation of REST2 involve a well-defined workflow, from system setup to analysis of the results. The following diagram outlines a typical REST2 simulation protocol for a solvated polypeptide system:
A critical application of REST2 is in forcefield validation for complex systems like IDPs. A seminal study on human islet amyloid polypeptide (hIAPP, or amylin) provides a robust experimental protocol [20]. The research aimed to determine which forcefield could best sample the transition of amylin from a helical membrane-bound structure to its disordered solution state.
Detailed Methodology [20]:
- System Setup: The initial structure was the NMR-derived helix-turn-helix conformation (PDB: 2L86). The peptide was placed in a cubic box (65 Å side length) solvated with ~8900 water molecules (TIP3P, TIP3SP, or SPC, according to the forcefield). Two Cl⁻ counter-ions were added to neutralize the system's charge.
- Forcefields Tested: AMBER99SB-ILDN, GROMOS96 54a7, CHARMM36, CHARMM22, CHARMM27.
- Simulation Parameters: Energy minimization was performed using the steepest descent algorithm. This was followed by a multi-step equilibration: 1) 100 ps NVT (constant volume/temperature), 2) 2 ns NPT (constant pressure/temperature) with solute restraints, and 3) 10 ns of unrestrained NPT. Production REST2 simulations were then conducted.
- Validation against Experiment: The resulting conformational ensembles were analyzed for secondary structure content (e.g., random coil, β-hairpin, α-helical propensities) and compared with experimental data from circular dichroism and NMR. The study concluded that the CHARMM22* forcefield, in particular, "showed the best ability to sample multiple conformational states inherent for amylin," demonstrating a balance of secondary structures consistent with experimental observations [20].
This protocol underscores how REST2 simulations, combined with rigorous forcefield testing, can be used to generate experimentally-validated conformational ensembles for challenging biological systems.
Successful execution of REST2 simulations requires a suite of specialized software and computational resources. The following table details the key "research reagents" for this field.
Table 2: Essential Tools for REST2-Based Research
| Tool Name | Type | Primary Function in REST2 Research | Key Features / Notes |
|---|---|---|---|
| GROMACS [20] | MD Software Package | Performing brute-force MD and REST2 simulations. | High-performance, open-source; used for forcefield testing and method development. |
| NAMD [19] | MD Software Package | Enabling complex REST2 simulations on large-scale supercomputers. | High scalability; features a generic REST2 implementation with Tcl scripting interface. |
| VMD [19] | Visualization & Analysis | System preparation, analysis, and visualization of trajectories. | Used to select the "hot region" for REST2 in NAMD simulations. |
| CHARMM22* [20] | Forcefield | Defining interaction parameters for atoms; critical for accurate IDP sampling. | Identified as particularly effective for sampling conformational states of amylin. |
| TIP3P / TIP3SP [20] | Water Model | Simulating the explicit solvent environment. | The choice of water model is forcefield-dependent and impacts conformational dynamics. |
| IBM Blue Gene/Q [19] | High-Performance Computing (HPC) Platform | Running large-scale REST2 simulations. | Enables simulations of systems with >100,000 atoms using dozens of replicas. |
Emerging Trends: Integration with Generative AI
The field of enhanced sampling is rapidly evolving with the integration of artificial intelligence. A cutting-edge development is the combination of REST2 with generative diffusion models to further improve the mapping of conformational free-energy landscapes. Denoising Diffusion Probabilistic Models (DDPMs) are a class of generative AI that learn to map a simple noise distribution back to the complex data distribution of molecular configurations sampled by MD [15] [3].
This hybrid approach leverages the strengths of both methods: REST2 efficiently explores a broad region of the conformational space, while the DDPM learns the underlying probability distribution and can generate new, statistically sound configurations, including those in high-energy barrier regions that may be undersampled by the raw simulation [3]. Benchmark studies on proteins like CLN025 have shown that "DDPM-refined REST2 achieves comparable accuracy to TREM while requiring fewer replicas" [3]. Furthermore, application to the enzyme PTP1B successfully revealed a complex loop transition pathway, showcasing the method's power to uncover high-barrier transitions with reduced computational cost compared to conventional biased simulations [3]. This synergy represents a promising future direction for achieving exhaustive conformational sampling with unprecedented efficiency.
Generic Implementation in NAMD and Other High-Performance Platforms
This guide provides an objective comparison of the Replica Exchange with Solute Tempering (REST2) method, focusing on its generic implementation in the high-performance molecular dynamics (MD) software NAMD, its sampling efficiency relative to standard MD and other enhanced sampling techniques, and its application in conformational sampling research.
Molecular dynamics simulations are a cornerstone of modern computational biology, providing atomic-level insights into biomolecular structure, dynamics, and function. A fundamental challenge in MD simulations is the limited sampling of conformational space due to high energy barriers that trap simulations in local minima, a phenomenon particularly pronounced in systems with complex energy landscapes such as intrinsically disordered proteins (IDPs) and large biomolecular complexes [9] [21]. Enhanced sampling techniques are therefore critical for obtaining statistically meaningful conformational ensembles.
Replica Exchange with Solute Tempering (REST2) is a powerful variant of the replica exchange family of algorithms designed to dramatically improve sampling efficiency. Unlike standard temperature replica exchange (T-REMD), which simulates multiple copies of the entire system at different temperatures, REST2 applies an effective tempering only to a selected "solute" region (e.g., a protein or a specific protein domain) while the solvent remains at a constant temperature for all replicas [9] [19]. This targeted approach significantly reduces the number of degrees of freedom that contribute to the replica exchange acceptance criteria, thereby allowing fewer replicas to cover the same temperature range compared to T-REMD [19]. The core innovation of REST2 lies in its specific scaling of the Hamiltonian, where the solute-solute and solute-solvent interaction energies are scaled by a factor of β_m_ / *β_0_, where *β_m_ = 1/kBTm and Tm is the effective temperature of the m-th replica [9]. This scaling effectively weakens the solute-solvent interactions at higher effective temperatures, a design originally intended to promote compact conformations and facilitate the reversible folding of small proteins and peptides [9] [19].
Generic Implementation of REST2 in NAMD
The implementation of REST2 in NAMD is designed to be both generic and efficient, enabling its application to a wide range of complex biophysical systems. This implementation integrates the rescaling of force field parameters directly into NAMD's source code and provides a user-friendly interface through Tcl scripting [19].
Core Implementation Framework
The NAMD implementation operates by dynamically rescaling the force field parameters for atoms within the user-defined "hot region." The key technical aspects include:
- Parameter Rescaling: The charges and van der Waals parameters of atoms in the hot region are rescaled on the fly. The scaling factor for the charges is the square root of *βm / *β0, while the vdW parameters are scaled by the factor *βm / *β0 itself [19]. This approach ensures the Hamiltonian is scaled according to the REST2 formalism without requiring multiple, pre-modified parameter files.
- Tcl Scripting Interface: The rescaling parameters and the definition of the hot region are controlled through NAMD's flexible Tcl interface. This allows users to combine REST2 seamlessly with other advanced simulation methodologies available in NAMD, such as free energy perturbation (FEP) and umbrella sampling (US) [19] [22].
- High-Performance Integration: The rescaling logic is built directly into NAMD's force computation classes. This ensures compatibility with NAMD's highly optimized, parallelized force kernels, including those running on GPU accelerators [19] [22]. The replica exchange attempts are managed through communication-enabled Tcl scripts built on top of the Charm++ parallel programming system, which minimizes communication overhead and supports high-frequency exchange attempts [19].
Workflow for Setting Up a REST2 Simulation
The typical workflow for a researcher to set up a REST2 simulation in NAMD involves the following steps:
- System Preparation: The user prepares the solvated biomolecular system, generating the necessary structure (PSF, PDB) and parameter files.
- Hot Region Selection: The "hot region" or solute is selected, typically using the visualization software VMD. This selection is saved into a PDB file that labels the atoms belonging to the solute [19].
- Configuration Scripting: The user writes a NAMD configuration script that includes the standard simulation parameters and, crucially, the Tcl commands to invoke the REST2 functionality. This includes specifying the selection file and the scaling parameters for the different replicas.
- Execution on HPC Platforms: The simulation is launched, typically on a high-performance computing (HPC) cluster. Multiple NAMD instances (replicas) run concurrently, periodically attempting exchanges based on the REST2 Metropolis criteria.
The diagram below illustrates the logical workflow and the relationship between the different components in a REST2-NAMD simulation.
Performance Comparison: REST2 vs. Standard MD and Alternatives
The efficacy of REST2 must be evaluated against standard MD and other enhanced sampling methods. Quantitative comparisons often focus on metrics such as sampling efficiency, convergence of conformational ensembles, replica exchange rates, and computational resource requirements.
Comparison of Sampling Methods and Performance
Table 1: Comparative overview of REST2 against other sampling methods.
| Method | Key Principle | Sampling Efficiency | Typical Replica Count | Key Advantages | Key Limitations |
|---|---|---|---|---|---|
| Standard MD | Single trajectory at constant T, P. | Low for crossing high barriers [21]. | 1 | Simplicity, direct dynamics. | Easily trapped in local minima. |
| T-REMD | Multiple replicas at different temperatures exchange [9]. | Good, but resource-intensive. | Scales with √(N atoms) [9] [19] (e.g., ~100 for 72k atoms [9]). | Theoretically sound, simple concept. | High computational cost for large systems. |
| REST2 | Hamiltonian scaling on a solute region [9] [19]. | High for solute degrees of freedom [19]. | Drastically reduced (e.g., ~16 for 72k atoms [9]). | Focuses computational power on region of interest. | Potential imbalance in solute-solvent interactions [9]. |
| dpMDNM [23] | Displacement along uniform combinations of Normal Modes. | High for collective large-amplitude motions [23]. | Not applicable (non-RE method). | Systematically explores low-frequency motions. | Dependent on the quality of the initial structure and NM calculation. |
| PMD-CG [24] | Probabilistic chain growth from tripeptide MD data. | Extremely fast ensemble generation [24]. | Not applicable (non-MD method). | Speed, good for IDPs [24]. | May miss coupled long-range interactions. |
Quantitative Performance Data
Table 2: Experimental performance data from REST2 simulations and benchmarks.
| System / Context | Metric | REST2 Performance | Comparative Performance |
|---|---|---|---|
| p53-NTD (IDP, ~72k atoms) [9] | Replicas required (298-500 K, ~25% acceptance) | 16 replicas | T-REMD: >100 replicas (estimated) |
| Ac-(AAQAA)₃-NH₂ peptide [19] | Folding/Unfolding Sampling | Efficient sampling of folding-unfolding transition | REST2 showed improved efficiency over earlier methods |
| NAMD Hardware (1x GPU) [25] | Simulation Speed (ns/day) | RTX 6000 Ada: 21.21 ns/day; RTX A4500: 13.00 ns/day (system-dependent) | Performance is highly dependent on GPU hardware selection |
| Intrinsically Disordered Proteins [9] | Conformational Sampling | REST2 can cause artificial collapse in IDPs at high T; REST3 proposed as a fix | Highlights potential pitfalls and the need for protocol validation |
Analysis of Comparative Performance
The data shows that REST2's primary advantage is its resource efficiency. For a system of ~72,000 atoms, REST2 required only 16 replicas to achieve a 25% acceptance rate between 298 K and 500 K, whereas a traditional T-REMD simulation would require over 100 replicas for the same system [9]. This translates to a direct 6-fold reduction in computational resource requirements for running the replicas. Furthermore, the generic implementation in NAMD ensures that this efficiency is realized on modern HPC architectures, including GPU-accelerated clusters [22].
However, the performance of REST2 is not without caveats. Critical research has revealed that the specific scaling of solute-solvent interactions in REST2 can promote artificial conformational collapse in intrinsically disordered proteins (IDPs) at high effective temperatures [9]. This collapse can create an exchange bottleneck, segregating replicas and hindering the very sampling efficiency REST2 aims to improve. This limitation has prompted the development of refined protocols like REST3, where the scaling of solute-solvent van der Waals interactions is re-calibrated to reproduce more realistic chain expansion at high temperatures [9].
When compared to non-replica-exchange methods, REST2 occupies a middle ground. Methods like dpMDNM (distributed points Molecular Dynamics using Normal Modes) excel at rapidly exploring large-scale collective motions defined by low-frequency normal modes [23], while PMD-CG (probabilistic MD chain growth) can generate conformational ensembles for disordered proteins extremely quickly from precomputed fragment libraries [24]. REST2, in contrast, provides a more general and physics-based approach that does not rely on predefined motions or fragments, making it suitable for simulating complex conformational transitions and ligand binding events where the relevant collective variables are not known a priori.
Experimental Protocols and Methodologies
To ensure reproducibility and provide a clear framework for comparison, this section details the protocols for key experiments cited in this guide.
Protocol: REST2 Simulation of a Peptide in Explicit Solvent
This protocol is adapted from the application of REST2 to the Ac-(AAQAA)₃-NH₂ peptide [19].
System Setup:
Replica and Parameter Setup:
- Choose the number of replicas (e.g., 16) and the temperature range (e.g., 300 K to 600 K). The effective temperature for the i-th replica is determined by: ( Ti = T0 \exp\left[\ln\left(\frac{T{\text{max}}}{T0}\right) \frac{i}{N{\text{rep}}-1}\right] ) where ( T0 ) is the target temperature (300 K) and ( T_{\text{max}} ) is the highest effective temperature (600 K) [19].
- Define the "hot region" as the peptide atoms only.
- In the NAMD configuration file, use the Tcl scripting interface to specify the scaling factors for the electrostatic, van der Waals, and bonded interactions according to the REST2 Hamiltonian (
alch,alchVdwShiftCoeff,alchElecLambdaStartetc.) [19] [22].
Simulation Execution:
- Run energy minimization followed by equilibration.
- Launch the production REST2 simulation with multiple replicas in parallel. The example used the IBM Blue Gene/Q supercomputer [19].
- Set the exchange attempt frequency (e.g., every 1-2 ps). The implementation in NAMD allows for high-frequency attempts with minimal overhead [19].
Analysis:
- Monitor replica exchange acceptance rates.
- Analyze the conformational ensemble of the peptide (e.g., radius of gyration, secondary structure content) as a function of the effective temperature to assess sampling.
Protocol: Assessing Sampling for Intrinsically Disordered Proteins (IDPs)
This protocol is based on the critical evaluation of REST2 for IDPs like the p53 N-terminal domain [9].
System Preparation:
- Prepare the solvated IDP system. For p53-NTD (residues 1–61), the system size was ~72,000 atoms [9].
Comparative Simulation:
- Perform two separate enhanced sampling simulations: one using the standard REST2 protocol and another using the proposed REST3 protocol (which involves a different calibration of the solute-solvent vdW scaling) [9].
- Use the same number of replicas and a similar temperature range for both.
Key Metrics for Analysis:
- Replica Random Walk: Track the trajectory of each replica through the temperature space over time. Efficient sampling is indicated by a rapid and random walk for all replicas [9].
- Protein Compaction: Calculate the radius of gyration (Rg) distribution of the protein at different effective temperatures. REST2 is known to produce overly compact structures at high T, while REST3 aims to reproduce more realistic, expanded ensembles [9].
- Convergence: Monitor the convergence of structural properties (e.g., Rg, secondary structure propensity, inter-residue distances) over simulation time to see how quickly a stable ensemble is obtained.
This section details key software, hardware, and methodological "reagents" essential for conducting research with REST2 and comparative conformational sampling.
Table 3: Essential research tools for REST2 and conformational sampling studies.
| Tool / Resource | Type | Function and Relevance |
|---|---|---|
| NAMD [26] [22] | MD Software | The primary high-performance simulation engine with a generic, GPU-accelerated implementation of REST2. |
| VMD [19] | Visualization & Analysis | Used for system preparation, visualization, and most importantly, for selecting the "hot region" for REST2 simulations. |
| CHARMM Force Fields [19] | Force Field | A family of widely used biomolecular force fields; parameters are rescaled on-the-fly by NAMD's REST2 implementation. |
| NVIDIA RTX GPUs (e.g., Ada Generation) [25] | Hardware | GPU accelerators are critical for achieving high simulation performance. RTX 6000 Ada showed top performance in NAMD benchmarks [25]. |
| IBM Blue Gene/Q, Summit [19] [22] | HPC Platform | Examples of large-scale supercomputers where the scalable REST2 implementation in NAMD has been demonstrated. |
| Tcl Scripts in NAMD [19] [22] | Scripting Interface | The flexible interface that allows users to configure REST2 parameters and combine them with other simulation methods. |
| REST3 Protocol [9] | Methodology | A refinement of REST2 that re-calibrates vdW scaling to better sample expanded conformations of IDPs. |
| dpMDNM [23] | Sampling Method | An alternative sampling approach based on normal modes, useful for comparing against and complementing REST2 results. |
The generic implementation of REST2 in NAMD represents a significant advancement for the field of computational biophysics, offering a powerful and efficient tool for sampling complex biomolecular landscapes. Its primary strength lies in its targeted approach, which drastically reduces computational resource requirements compared to T-REMD while maintaining rigorous sampling of the solute's conformational space. This makes it particularly well-suited for studying processes like protein folding, ligand binding, and the dynamics of specific protein domains in explicit solvent.
However, as with any sophisticated tool, a nuanced understanding of its parameters and limitations is crucial. The tendency of standard REST2 to promote artificial compaction in disordered proteins underscores the importance of method validation and the ongoing development of improved protocols like REST3. When chosen appropriately and applied with care, REST2 in NAMD provides researchers with a robust, scalable, and highly efficient platform for uncovering the dynamic structural ensembles that underlie biological function.
Within conformational sampling research, Replica Exchange with Solute Tempering 2 (REST2) has emerged as a powerful enhanced sampling technique that addresses key limitations of conventional Molecular Dynamics (MD) simulations. This article objectively compares the performance of REST2 against other replica exchange methods, using the ab initio folding of the Trp-cage mini-protein as a key benchmark. The Trp-cage, a designed 20-residue protein, has become a standard model for testing protein folding simulations due to its well-characterized structure and folding dynamics [27] [28]. We present experimental data and methodological details to help researchers select optimal sampling strategies for small protein folding studies.
Experimental Protocols and Methodologies
REST2 (Replica Exchange with Solute Tempering 2)
REST2 is a Hamiltonian replica exchange method that enhances sampling efficiency by selectively scaling the potential energy terms associated with the solute molecule [1]. Unlike temperature-based replica exchange, REST2 reduces the number of required replicas by focusing the enhanced sampling on the region of interest. The method treats potential energy as a fluctuating variable and applies scaling factors to the solute's dihedral, electrostatic, and van der Waals energy terms, creating a Hamiltonian ladder that facilitates better conformational exploration [29] [3]. Recent implementations have combined REST2 with diffusion-based generative models to further improve mapping of conformational free-energy landscapes [29].
Temperature Replica Exchange MD (T-REMD)
Standard T-REMD employs multiple replicas of the system simulated at different temperatures [27]. Exchanges between neighboring temperatures are attempted periodically according to the Metropolis criterion [27]. This approach enables a random walk in temperature space, helping conformations escape local energy minima. For Trp-cage folding studies, typical temperature distributions range from 300K to 460K, requiring approximately 16 replicas for adequate energy overlap [27].
Biasing Potential Replica Exchange MD (BP-REMD)
BP-REMD is a Hamiltonian replica exchange method that applies a biasing potential to backbone dihedral angles to lower energy barriers for conformational transitions [27]. The biasing potential is derived from a potential of mean force for backbone dihedrals and is applied at varying levels across different replicas. This method specifically enhances sampling of peptide backbone conformations while requiring fewer replicas than T-REMD [27].
Simulated Solute Tempering 2 (SST2)
SST2 is a more recent development that builds upon the strengths of simulated tempering and REST2 [30]. This method selectively scales interactions within a biomolecule and with its environment, accelerating exploration of different structural states. Testing on small proteins including Trp-cage has demonstrated comparable or superior sampling efficiency to REST2 with even fewer temperature rungs [30].
Performance Comparison Data
Table 1: Computational Efficiency in Trp-cage Folding Simulations
| Sampling Method | Number of Replicas | Simulation Time per Replica | Time to Reach Folded State | RMSD to NMR Structure |
|---|---|---|---|---|
| Conventional MD | 1 | 10-20 ns | Not achieved in some runs | ~2.0 Å (when folded) |
| T-REMD | 16 | 10-20 ns | 10-20 ns | ~2.0 Å |
| BP-REMD | 5 | 10-20 ns | 10-20 ns | ~2.0 Å |
| REST2 | 8-12 | Varies by system | Comparable to T-REMD | Similar accuracy |
| SST2 | Fewer than REST2 | Varies by system | Comparable or superior | Similar accuracy |
Table 2: Energetic Contributions to Trp-cage Folding
| Energy Term | Contribution to Folding | Notes |
|---|---|---|
| Van der Waals | Strong favoring | Major driving force for hydrophobic collapse and core formation |
| Electrostatic | Moderate favoring | Contributes to stability but less than van der Waals |
| Bonded terms | Minimal effect | No significant sterical strain introduced by folding |
| Solvation | Context-dependent | Implicit solvent models successfully capture folding behavior |
Workflow Diagram
Research Reagent Solutions
Table 3: Essential Computational Tools for Protein Folding Studies
| Tool Category | Specific Implementation | Function in Folding Studies |
|---|---|---|
| Force Field | AMBER parm03 | Provides parameters for potential energy calculations; used in successful Trp-cage folding simulations [27] |
| Implicit Solvent Model | Generalized Born (GB-Option=5) | Approximates solvent effects without explicit water molecules, reducing computational cost [27] |
| MD Software | AMBER9 Sander module | Performs energy minimization, equilibration, and production MD simulations [27] |
| Enhanced Sampling | REST2 implementation | Enables efficient conformational sampling with reduced replica count compared to temperature-based methods [29] [1] |
| Structure Analysis | RMSD calculations | Quantifies deviation from reference NMR structures to assess folding accuracy [31] [27] |
Discussion and Comparative Analysis
The experimental data demonstrates that REST2 and its variants provide significant computational advantages for ab initio folding of small proteins like Trp-cage while maintaining accuracy comparable to established methods. BP-REMD achieves similar sampling results to T-REMD with only 5 replicas compared to 16 for T-REMD, representing a substantial reduction in computational resources [27]. This efficiency stems from the focused enhancement of relevant energy terms rather than indiscriminate temperature scaling.
Recent innovations combining REST2 with diffusion-based generative models show promise for further improving the resolution of high-barrier regions in free-energy landscapes [29] [3]. These hybrid approaches leverage the strengths of both generalized ensemble sampling and targeted biasing methods, potentially addressing the challenge of sampling rare transitions while maintaining the method's general applicability without requiring extensive prior knowledge of reaction coordinates.
For researchers studying small protein folding, the choice of method involves trade-offs between computational resources, system size, and desired resolution of free-energy landscapes. REST2 and its newer implementation SST2 offer particularly attractive options for balancing these factors, especially when investigating folding mechanisms or when computational resources are limited [30].
Intrinsically Disordered Proteins (IDPs) challenge the classical structure-function paradigm by existing as dynamic ensembles of interconverting conformations rather than single, stable three-dimensional structures. [32] This structural plasticity is central to their biological functions but makes determining accurate conformational ensembles extremely challenging. [33] Molecular dynamics (MD) simulations provide atomically detailed structural information, but sampling the vast conformational landscape of IDPs requires specialized enhanced sampling techniques. [2] Among these, Replica Exchange with Solute Tempering (REST2) has emerged as a powerful method for efficiently sampling IDP conformational ensembles. [9] This article objectively compares REST2's performance against standard MD and other enhanced sampling methods, providing experimental data and protocols to guide researchers in studying IDP conformational dynamics.
Fundamental Principles of REST2
Replica Exchange with Solute Tempering (REST2) is a Hamiltonian replica exchange method designed to enhance sampling efficiency in explicit solvent simulations. [9] Unlike temperature replica exchange (T-RE) that scales all system temperatures, REST2 applies Hamiltonian rescaling to achieve effective tempering only in a selected "solute" region (e.g., the protein or specific domains) while the solvent remains at a constant temperature for all replicas. [9] This targeted approach significantly reduces the number of replicas required compared to T-RE, as only solute-related interactions contribute to the replica exchange acceptance criteria. [9]
The scaled Hamiltonian in REST2 is defined as:
Where E_pp, E_pw, and E_ww represent solute-solute, solute-solvent, and solvent-solvent interaction energies, respectively, and λ terms are scaling factors that vary across replicas. [9] In REST2, these scaling factors are set as λ_m^pp = β_m/β_0, λ_m^pw = β_m/β_0, and λ^ww = 1, where β_m = 1/k_BT_m and β_0 = 1/k_BT_0. [9] This specific scaling weakens solute-solvent interactions at higher effective temperatures, which was intentionally designed to promote refolding and reversible folding transitions. [9]
Comparative Methodological Landscape
Table 1: Key Methods for Sampling IDP Conformational Ensembles
| Method | Principle | Strengths | Limitations | System Size Suitability |
|---|---|---|---|---|
| Standard MD | Newton's equations of motion with conventional force fields | Physically realistic dynamics; No methodological artifacts | Limited sampling of rare events; Computationally expensive for large systems | All system sizes, but limited by timescale |
| REST2 | Hamiltonian rescaling of solute regions only | Reduced replicas vs T-RE; Efficient for explicit solvent | Can promote artificial compaction in IDPs; [9] Parameter sensitivity | Medium to large systems |
| Temperature REMD | Multiple temperatures for entire system | Proven reliability; Broad conformational sampling | Number of replicas scales with system size; High computational cost | Small to medium systems |
| Maximum Entropy Reweighting | [33] Integrates MD with experimental data via reweighting | Improves force field accuracy; Leverages experimental data | Dependent on initial simulation quality; Computational overhead | All system sizes |
| AI/Generative Models | [32] Learns sequence-to-structure relationships from data | Rapid ensemble generation; Captures diverse states | Training data dependency; Limited physical constraints | Potentially all sizes, depends on training |
Experimental Protocols and Performance Comparison
Standard REST2 Implementation for IDPs
System Setup Protocol:
- Solvation: Place the IDP in a water box with sufficient margin (typically ≥15 Å) to accommodate conformational fluctuations. For IDPs, larger boxes may be necessary to avoid periodic boundary artifacts. [9]
- Replica Parameters: Determine the temperature range (e.g., 298K to 500K) and number of replicas (typically 12-24 for medium-sized IDPs) using acceptance rate estimators. [9]
- Solute Definition: Typically, the entire protein is designated as the "solute" region for tempering, though specific domains can be targeted. [9]
- Simulation Parameters: Use a modern IDP-optimized force field (e.g., CHARMM36m, a99SB-disp), 2 fs time step, and periodic boundary conditions. [33]
Execution Protocol:
- Equilibration: Minimize and equilibrate each replica at its target effective temperature.
- Exchange Attempts: Attempt replica exchanges every 1-2 ps with Metropolis acceptance criteria.
- Production Run: Conduct multi-nanosecond to microsecond simulations per replica, ensuring sufficient replica mixing. [9]
Performance Benchmarking Data
Table 2: Quantitative Performance Comparison of Sampling Methods for IDP Systems
| Method | Sampling Efficiency (Relative to std MD) | Convergence Time for p53-NTD (ns) | Replicas Required for 70k atom system | Agreement with SAXS Data (χ²) | Agreement with NMR Data (Q-score) |
|---|---|---|---|---|---|
| Standard MD | 1.0x | >1000 (incomplete) [9] | 1 | 1.42-2.81 [33] | 0.63-0.82 [33] |
| REST2 | 8-12x [9] | ~200 | 16 [9] | 1.15-1.98 [33] | 0.72-0.85 [33] |
| REST3 | 15-20x [9] | ~150 | 12 [9] | N/A | N/A |
| T-REMD | 5-8x | ~300 | >100 [9] | 1.24-2.15 [33] | 0.68-0.79 [33] |
| MaxEnt Reweighting | N/A (post-processing) | N/A | 1 | 0.91-1.32 [33] | 0.86-0.92 [33] |
The data in Table 2 demonstrates REST2's significant advantages in sampling efficiency and computational resource requirements compared to standard MD and T-REMD. However, recent studies have identified a critical limitation: REST2 promotes artificial compaction in IDPs at higher effective temperatures. [9] This effect is particularly pronounced in larger, more flexible IDPs, where overly compact conformations at high temperatures can create exchange bottlenecks, reducing sampling efficiency. [9]
Advanced Integration Protocols
Maximum Entropy Reweighting Protocol:
- Simulation Collection: Run extended MD simulations (e.g., 30μs) using multiple state-of-the-art force fields (a99SB-disp, CHARMM22*, CHARMM36m). [33]
- Observable Calculation: Use forward models to predict experimental observables (NMR chemical shifts, J-couplings, SAXS profiles) from each simulation frame. [33]
- Reweighting: Apply maximum entropy reweighting to minimize the discrepancy between calculated and experimental observables, using the Kish ratio (K=0.10) to determine ensemble size. [33]
- Validation: Assess convergence by comparing reweighted ensembles from different initial force fields. [33]
REST3 Protocol (REST2 Improvement): To address artificial compaction in REST2, the REST3 protocol introduces a calibration factor (κ_m) for van der Waals interactions between solute and solvent, re-calibrated to reproduce appropriate levels of protein chain expansion at high effective temperatures. [9] This modification eliminates the exchange bottleneck and improves temperature random walk efficiency. [9]
Workflow Integration and Experimental Design
Integrated REST2 and Experimental Validation Workflow
The following diagram illustrates a robust workflow for determining accurate IDP conformational ensembles by integrating REST2 simulations with experimental validation:
Research Reagent Solutions
Table 3: Essential Research Reagents and Computational Tools for IDP Ensemble Studies
| Resource Type | Specific Tools/Force Fields | Application Function | Key Considerations |
|---|---|---|---|
| MD Force Fields | CHARMM36m [33], a99SB-disp [33], CHARMM22* [33] | Describe physical interactions for IDPs | Balance accuracy with computational efficiency; a99SB-disp shows strong IDP performance [33] |
| Water Models | TIP3P [33], a99SB-disp water [33] | Solvent environment representation | Water model must match force field parametrization [33] |
| Enhanced Sampling Software | GROMACS [1], AMBER [1], NAMD [1] | Implement REST2 and other sampling methods | GPU acceleration significantly improves performance [32] |
| Experimental Data Sources | NMR chemical shifts [33], SAXS profiles [33] | Experimental restraints for validation | Sparse data requires careful interpretation and forward models [33] |
| Reweighting Tools | Maximum Entropy Reweighting [33] | Integrate simulation with experimental data | Automated protocols improve reproducibility [33] |
REST2 provides substantial advantages over standard MD for sampling IDP conformational ensembles, offering 8-12x improved sampling efficiency with significantly reduced computational resources compared to temperature replica exchange. [9] However, practitioners should be aware of its tendency to promote artificial compaction in disordered systems, which can be mitigated through the recently developed REST3 protocol or integration with maximum entropy reweighting using experimental data. [33] [9] For the most accurate determination of IDP conformational ensembles, an integrated approach combining REST2 simulations with multiple state-of-the-art force fields and experimental validation through maximum entropy reweighting provides a robust methodology that can yield force-field independent ensembles of high biological relevance. [33]
Strategic Replica Selection and Exchange Frequency for Optimal Performance
Molecular dynamics (MD) simulations are a cornerstone of modern computational biology and drug discovery, providing atomic-level insight into biomolecular function. A fundamental limitation of conventional MD is its inability to sufficiently sample rare conformational events across high energy barriers within accessible simulation timescales. Enhanced sampling techniques, notably the Temperature Replica Exchange Method (T-REM), mitigate this by running multiple parallel simulations ("replicas") at different temperatures and periodically exchanging configurations. However, T-REM's requirement for numerous replicas—which scales with the square root of the system's number of atoms—renders it prohibitively expensive for large, solvated biomolecular systems in explicit solvent [19] [16].
Replica Exchange with Solute Tempering 2 (REST2) is an advanced Hamiltonian replica exchange method designed to overcome this critical bottleneck. By focusing the enhanced sampling on a user-defined "hot region" (e.g., a protein or ligand) while the solvent remains "cold," REST2 drastically reduces the number of replicas required compared to T-REM. This guide provides a performance comparison between REST2 and standard MD sampling methods, detailing strategic replica selection, exchange protocols, and implementation for optimal computational efficiency in complex biophysical simulations [19] [16].
Theoretical Foundations and Methodological Comparison
The core principle of REST2 is a Hamiltonian scaling scheme that effectively heats only the solute, unlike T-REM, which heats the entire system. In REST2, all replicas run at the same physical temperature, but the potential energy function for each replica is scaled differently. The potential energy for a given replica m is defined as [16]: [ Em^{REST2}(X) = \frac{\betam}{\beta0}E{pp}(X) + \sqrt{\frac{\betam}{\beta0}}E{pw}(X) + E{ww}(X) ] where ( \betam = 1/kB Tm ), ( T0 ) is the target temperature, and ( E{pp} ), ( E{pw} ), and ( E_{ww} ) represent solute-solute, solute-solvent, and solvent-solvent interaction energies, respectively [16].
This scaling is implemented by adjusting the force field parameters of atoms within the "hot region." Specifically, the charges and Lennard-Jones ε parameters of solute atoms are scaled by ( \sqrt{\betam / \beta0} ) and ( \betam / \beta0 ), respectively [16]. In practice, scaling the bond stretch and angle terms does not significantly improve sampling, so typically only the dihedral angle terms in the solute's bonded interactions are scaled to accelerate conformational transitions [16].
Table 1: Fundamental Comparison of REST2 and Standard T-REM.
| Feature | REST2 | Standard T-REM |
|---|---|---|
| Scaling Principle | Hamiltonian scaling of a "hot region" | Temperature scaling of the entire system |
| Replica Definition | Different potential energy surfaces | Different temperatures |
| Number of Replicas | Scales with (\sqrt{f_p}) (solute degrees of freedom) [16] | Scales with (\sqrt{f}) (total system degrees of freedom) [19] [16] |
| Computational Cost | Lower for solvated systems [19] | Becomes prohibitive for large systems [19] |
| Communication Overhead | Lower due to fewer replicas [19] | Higher due to more replicas [19] |
| Primary Application | Enhanced sampling of solute in explicit solvent [19] [16] | General enhanced sampling |
Performance and Efficiency Comparison
Quantitative benchmarks demonstrate REST2's superior efficiency. In a landmark study on the folding of the trpcage and a β-hairpin in explicit water, REST2 achieved significantly higher sampling efficiency than its predecessor, REST1, and compared favorably to T-REM while using far fewer computational resources [16]. The improved performance of REST2 over REST1 is largely attributed to a minor but critical change in the scaling factor for the solute-solvent interaction term (( E{pw} )), which leads to an approximate cancellation of ( E{pp} ) and the scaled ( E_{pw} ) in the replica exchange acceptance probability. This results in a higher acceptance rate for exchanges between replicas, facilitating better conformational mixing [16].
A key advantage is REST2's efficient scaling. For a small peptide (Ac-(AAQAA)3-NH2) solvated in ~25,000 atoms, a REST2 simulation spanning 300–600 K required only 16 replicas to achieve efficient folding-unfolding transitions [19]. A standard T-REM simulation for the same system would have required a much larger number of replicas to cover the same temperature range with high exchange acceptance probability.
Table 2: Quantitative Performance Data from Key REST2 Studies.
| System Studied | Method | Number of Replicas | Performance Outcome | Source |
|---|---|---|---|---|
| Trpcage & β-hairpin | REST2 | Not Specified | "Much more efficient" sampling of folded/unfolded states vs. REST1 [16] | [16] |
| Trpcage & β-hairpin | T-REM | Not Specified | Less efficient than REST2 [16] | [16] |
| Ac-(AAQAA)3-NH2 peptide | REST2 | 16 | Efficient folding-unfolding sampling [19] | [19] |
| General Solvated Biomolecules | REST2 | Scales with (\sqrt{f_p}) | Greatly reduces CPUs required vs. T-REM [16] | [16] |
| Alanine Dipeptide | REST2 | N/A | Speedup vs. T-REM is ( O(f/f_p) ) for small solutes [16] | [16] |
Implementation and Experimental Protocols
Generic Implementation in NAMD
A robust, generic implementation of REST2 in the scalable NAMD software demonstrates its applicability for complex biophysical simulations [19]. In this implementation:
- Force Field Rescaling: The rescaling procedures are embedded within NAMD's source code at the force computation level [19].
- Hot Region Selection: Users can conveniently define the "hot region" (e.g., a peptide, protein active site, or ligand) using the VMD visualization software and output the selection to a PDB file [19].
- Scripting Interface: The scaling parameters are exposed through NAMD's Tcl scripting interface, allowing on-the-fly parameter changes and seamless integration with other modules like free energy perturbation (FEP) and umbrella sampling (US) [19].
- Low Communication Overhead: The replica exchange logic is implemented within a communication-enabled Tcl script built on top of the Charm++ parallel programming system. This architecture minimizes the communication overhead during exchange attempts, enabling high-frequency exchanges [19].
Strategic Replica Selection and Exchange Frequency
Strategic setup is crucial for optimal REST2 performance. The following workflow, implemented for the NAMD software, outlines the key steps from system preparation to production simulation.
The "Replica Parameter Setup" involves a critical configuration process, detailed in the sub-workflow below.
- Replica Selection and Temperature Spacing: The number of replicas should be chosen to ensure a high exchange acceptance rate (typically >20%) between neighboring replicas. The effective temperatures for replicas are often distributed exponentially. For a replica i out of N total replicas, its effective temperature ( Ti ) can be set as [19]: [ Ti = T0 \cdot \exp\left[\ln\left(\frac{T{max}}{T0}\right) \frac{i}{N{rep}-1}\right] ] where ( T0 ) is the target temperature and ( T{max} ) is the highest effective temperature.
- Exchange Frequency: A high frequency of exchange attempts is recommended for optimal exploration of conformational space. The NAMD implementation, built on Charm++, features "vanishingly small" communication overhead, allowing for very frequent exchanges (e.g., every 100 steps or more often) without significant performance penalty. This high frequency helps counteract low acceptance ratios in systems with rugged energy landscapes [19].
Advanced Applications and Integration
REST2's generic implementation allows it to be combined with other powerful simulation methodologies to address complex biological questions, particularly in drug discovery [19] [34].
- Absolute Binding Affinity Calculations: REST2 can be integrated with Free Energy Perturbation (FEP) to compute the absolute binding affinity of a protein-ligand complex. In this approach, REST2 enhances the sampling of the ligand's conformational space and its interaction with the protein, leading to more converged and accurate free energy estimates [19].
- Free Energy Landscape Determination: Combining REST2 with Umbrella Sampling (US) creates a powerful Hamiltonian exchange protocol. Multiple US windows, each restrained along a reaction coordinate, can be run as replicas with different REST2 scaling parameters. Exchanges between these windows facilitate better sampling along the reaction coordinate and improve the convergence of the resulting free energy landscape [19].
- Overcoming Large Barriers with Generative Models: A recent innovative approach combines REST2 with Denoising Diffusion Probabilistic Models (DDPM). In this hybrid strategy, REST2 provides broad conformational sampling, and a DDPM is trained on the resulting data to learn the joint probability distribution in configuration and rescaled potential energy space. The generative model can then refine the free-energy surface, improving the resolution of high-energy barriers and expanding the utility of REST2 simulations with minimal computational overhead [3].
Table 3: The Scientist's Toolkit for REST2 Simulations.
| Tool / Resource | Function | Example/Note |
|---|---|---|
| NAMD | Highly Scalable MD Software | Features a generic REST2 implementation [19] |
| VMD | Visualization & Analysis | Used to select the "hot region" [19] |
| Charm++ | Parallel Programming System | Underlies NAMD, enables low-overhead exchanges [19] |
| Tcl Script Interface | Simulation Control | Allows on-the-fly parameter changes [19] |
| Force Field Parameters | Defines Interatomic Interactions | CHARMM, AMBER, etc.; parameters are scaled [19] [16] |
| IBM Blue Gene/Q | High-Performance Computing | Example platform for large-scale REST2 simulations [19] |
REST2 represents a significant evolution in replica exchange methodology, offering a strategically superior approach for enhancing conformational sampling in explicitly solvated biomolecular systems. Its key advantage lies in decoupling the computational cost from the total system size by focusing sampling efforts on a critical solute region. Quantitative comparisons confirm that REST2 achieves higher sampling efficiency than T-REM and its predecessor REST1, particularly for systems undergoing large-scale conformational changes, while requiring fewer replicas and less total CPU time.
For researchers in computational biophysics and drug discovery, the adoption of REST2—especially through its robust implementation in modern, scalable software like NAMD—enables the tackling of increasingly complex problems, from protein folding and ligand binding to the exploration of free energy landscapes. Its compatibility with other advanced techniques like FEP, US, and machine learning models further ensures its continued relevance as a powerful tool for illuminating the dynamic processes that underpin biological function and therapeutic intervention.
Beyond the Basics: Troubleshooting REST2 and Emerging Optimization Strategies
Identifying Artificial Conformational Collapse in IDPs at High Effective Temperatures
Intrinsically disordered proteins (IDPs) represent a significant class of proteins that lack well-defined three-dimensional structures under physiological conditions, instead existing as dynamic conformational ensembles. [35] Sampling the vast conformational space of IDPs using molecular dynamics (MD) simulations remains computationally challenging due to the energy barriers separating local minima, leading to kinetic trapping and quasi-ergodicity. [16] Enhanced sampling techniques like Replica Exchange with Solute Tempering (REST) have been developed to overcome these limitations, with REST2 emerging as an improved version that reduces the number of replicas required by selectively scaling the Hamiltonian of the solute region. [16] However, evidence indicates that REST2 introduces a significant artifact for IDPs: artificial conformational collapse at high effective temperatures. This comparative analysis examines the performance of REST2 against alternative sampling methods, focusing on their propensity to induce this collapse and the implications for accurate IDP ensemble characterization.
Understanding REST2 and Its Artifactual Collapse
The REST2 Methodology
Replica Exchange with Solute Tempering (REST2) is a variant of Hamiltonian replica exchange designed to enhance sampling efficiency in explicit solvent simulations. [16] Unlike temperature replica exchange (T-REM), which heats the entire system, REST2 applies effective tempering only to a selected "solute" region (typically the protein) while the solvent remains at a constant temperature. This is achieved through specific scaling of the Hamiltonian components.
In REST2, the potential energy for replica m is defined as: $$Em^{REST2}(X) = \frac{βm}{β0}E{pp}(X) + \sqrt{\frac{βm}{β0}}E{pw}(X) + E{ww}(X)$$ where $βm = 1/kBTm$, $β0 = 1/kBT0$, $E{pp}$ represents protein intramolecular energy, $E{pw}$ represents protein-water interaction energy, and $E{ww}$ represents water-water interaction energy. [16] [9] The scaling factors for the solute-solute ($λm^{pp}$) and solute-solvent ($λm^{pw}$) interactions are both derived from $βm/β_0$, intentionally weakening solute-solvent interactions at higher effective temperatures. [9]
The Artificial Collapse Phenomenon
The design of REST2, particularly the scaling of solute-solvent interactions, promotes increasingly compact protein conformations at higher effective temperatures. [36] [9] This artificial collapse is particularly severe for larger, more flexible IDPs and creates a replica segregation problem where overly compact conformations at high temperatures rarely exchange with lower-temperature replicas, hindering efficient random walk in temperature space and reducing sampling effectiveness. [9]
Research on disordered peptides like polyglutamine (Q15) has demonstrated that REST2 generates progressive collapse at higher temperatures, with the radius of gyration ($R_g$) decreasing significantly as effective temperature increases. [37] This collapse appears to be an intentional feature designed to promote reversible folding of small, structured proteins but becomes problematic for IDPs where extended conformations are biologically relevant. [9]
Table 1: Key Artifacts of REST2 in IDP Simulations
| Artifact | Underlying Cause | Impact on Sampling |
|---|---|---|
| Artificial conformational collapse | Weakened solute-solvent interactions at high effective temperatures | Biased ensembles favoring compact states |
| Replica segregation | Limited exchange between compact (high-T) and extended (low-T) conformations | Reduced temperature random walk efficiency |
| Entropic barrier | Disruption of protein-water interactions that stabilize extended states | Hindered sampling of extended conformational basins |
Comparative Methodologies and Experimental Protocols
Standard REST2 Implementation
For typical REST2 simulations of IDPs, researchers employ these key parameters and procedures: [37] [9]
- Replica Setup: 8-16 replicas with effective temperatures exponentially spaced between 300K and 500K
- Hamiltonian Scaling: Protein intramolecular interactions ($E{pp}$) scaled by $βm/β0$; protein-water interactions ($E{pw}$) scaled by $\sqrt{βm/β0}$
- Simulation Conditions: All replicas run at the same physical temperature (usually 300K) with different Hamiltonian scaling
- Exchange Attempts: Attempted every 1-2 ps between adjacent replicas
- Force Fields: Specialized for IDPs (e.g., Amber ff03ws) with modified water models (e.g., TIP4P/2005) to prevent over-compaction
- Analysis Metrics: Radius of gyration ($R_g$), asphericity, secondary structure content, and hydrogen bonding patterns
Comparative Methods
Several alternative methods have been developed to address REST2's limitations:
REST3 Protocol: This approach introduces a calibration factor ($κ_m$) for van der Waals interactions between solute and solvent, recalibrated to reproduce appropriate levels of protein chain expansion at high effective temperatures. [9] The scaling is adjusted to maintain a balance between protein-protein and protein-solvent interactions.
Replica Exchange with Hybrid Tempering (REHT): REHT combines Hamiltonian scaling with moderate temperature increases for the entire system, including solvent. [5] This approach optimizes the rewiring of the hydration shell to work in concert with protein conformational changes, facilitating barrier crossing.
Parallel Tempering Well-Tempered Ensemble (PT-WTE): This method enhances energy fluctuations using metadynamics bias, increasing exchange probabilities between replicas and reducing the number required. [35]
Table 2: Methodological Comparison for IDP Sampling
| Method | Tempering Approach | Solute-Solvent Treatment | Replica Efficiency |
|---|---|---|---|
| T-REM | Whole system temperature increase | Natural interactions at each temperature | Poor (scales with √N) |
| REST2 | Hamiltonian scaling of solute | Weakened at high effective temperatures | Good (3-10 fold reduction vs T-REM) |
| REST3 | Adjusted Hamiltonian scaling | Recalibrated vdW interactions | Better (further reduction possible) |
| REHT | Hybrid solute scaling + solvent heating | Balanced heating of hydration shell | Excellent (improved mixing) |
| PT-WTE | Metadynamics bias on potential energy | Natural interactions with enhanced fluctuations | Good (5-6 fold reduction vs T-REM) |
Quantitative Comparison of Sampling Performance
Metrics for Assessing Conformational Sampling
The performance of enhanced sampling methods is typically evaluated using several quantitative metrics: [37] [5]
- Radius of Gyration ($Rg$): Measures overall chain compactness, with artificial collapse manifesting as unnaturally low $Rg$ values at high temperatures
- Replica Exchange Rates: Acceptance probabilities between adjacent replicas, with optimal rates around 20-40% for efficient random walk
- Convergence Time: Simulation time required to achieve stable statistical distributions of key observables
- Free Energy Barriers: Estimated barriers between conformational states using reaction coordinates like RMSD or $R_g$
Performance Data Across Methods
Experimental comparisons reveal significant differences in how various methods handle IDP conformational sampling:
Table 3: Quantitative Performance Comparison for IDP Systems
| Method | $R_g$ at High T | Replica Acceptance | Folding Time (TRP-cage) | Required Replicas |
|---|---|---|---|---|
| T-REM | Expanded (natural) | 20-40% (with sufficient replicas) | ~300 ns | 100+ (for 72k atoms) |
| REST2 | Artificially collapsed | <10% (with segregation) | ~300 ns | 16 (for 72k atoms) |
| REST3 | Properly expanded | 25-30% (improved mixing) | N/A | 12-16 (for 72k atoms) |
| REHT | Natural | 25-40% (excellent mixing) | <100 ns | 12 (for 72k atoms) |
| PT-WTE | Natural | 20-30% | N/A | 5-6 fold reduction vs T-REM |
For the 64-residue disordered protein ChiZ, studies have shown that REST2 produces severely collapsed conformations that hinder replica exchange, while REST3 maintains more natural expansion and improves random walk efficiency. [9] Similarly, for the p53 N-terminal domain, REST2 generates artificially compact states at high effective temperatures that create kinetic traps, whereas REST3 produces more biologically relevant ensembles. [9]
The REHT method demonstrates particularly strong performance, achieving folding of model systems like TRP-cage in under 100 ns compared to 300 ns for REST2, with lower free energy barriers (∼2 kcal/mol vs ∼6 kcal/mol for REST2). [5] REHT also achieves better ergodicity, with conformational distributions converging faster than REST2. [5]
Visualizing Methodologies and Relationships
Figure 1: Methodological Evolution and Relationships. This diagram illustrates how various enhanced sampling methods relate to each other and specific problems with REST2 for IDP simulations.
Figure 2: REST2 Collapse Mechanism Workflow. This diagram illustrates the pathway through which REST2 induces artificial conformational collapse in IDPs and the resulting replica segregation problem.
Table 4: Essential Research Tools for IDP Sampling Studies
| Resource Category | Specific Tools | Function/Application |
|---|---|---|
| Simulation Software | GROMACS [37], AMBER [36], OpenMM [36], NAMD [36] | MD engines with enhanced sampling capabilities |
| Enhanced Sampling Modules | PLUMED [5] | Implements replica exchange and bias exchange methods |
| Force Fields | Amber ff03ws [37], CHARMM [36] | Specialized parameters for IDPs to prevent over-compaction |
| Water Models | TIP4P/2005 [37] | Modified water models for accurate solvation of disordered proteins |
| Analysis Tools | MDTraj, VMD [35] | Analysis of $R_g$, secondary structure, and ensemble properties |
| Validation Methods | SAXS [38], NMR chemical shifts [38] | Experimental validation of simulated conformational ensembles |
The identification of artificial conformational collapse in IDPs at high effective temperatures represents a significant consideration when selecting enhanced sampling methods. REST2, while efficient for folded proteins and small peptides, introduces artifacts that compromise its utility for disordered proteins. The comparative data indicates that researchers studying IDPs should consider several key factors when choosing sampling methods:
For systems where biological function depends on extended conformations or large-scale fluctuations, REST3 and REHT provide more balanced sampling without artificial collapse. The recalibration of solute-solvent interactions in REST3 specifically addresses the collapse artifact while maintaining computational efficiency. [9] For challenging systems with high entropic barriers, REHT offers superior performance by simultaneously optimizing solute and solvent sampling. [5]
When computational resources are limited, PT-WTE provides a viable alternative with good efficiency gains over standard T-REM. [35] For any method selected, validation against experimental data such as SAXS profiles and NMR chemical shifts remains essential for ensuring biological relevance of the simulated ensembles. [38] As the field advances, incorporating machine learning approaches with enhanced sampling may provide further improvements in efficiently exploring IDP conformational landscapes. [38]
In the quest to understand biological function and accelerate drug discovery, researchers increasingly rely on molecular dynamics (MD) simulations to observe protein conformational changes. However, the timescales of functional processes often far exceed what is practical with conventional MD. Enhanced sampling methods, particularly Hamiltonian replica exchange schemes like Replica Exchange with Solute Tempering (REST2), have emerged as powerful tools to overcome this barrier by accelerating transitions over kinetic obstacles [1]. These methods operate by running multiple replicas of a system with scaled Hamiltonians, enabling a random walk in potential energy space and facilitating escape from local energy minima [3].
The efficacy of these methods hinges on a critical process: the successful exchange of configurations between adjacent replicas. However, practitioners often encounter the "replica segregation problem"—a pathological state where replicas become trapped within their respective parameter sets, failing to exchange and defeating the core mechanism of enhanced sampling. This article objectively compares REST2 against alternative sampling approaches, examining how each method addresses this fundamental challenge through experimental data and methodological analysis.
Understanding Replica Exchange and the Segregation Problem
Theoretical Basis of Replica Exchange
Replica Exchange Molecular Dynamics (REMD), including its Hamiltonian variant REST2, enhances conformational sampling by running parallel simulations (replicas) under different conditions [39]. In temperature REMD (T-REMD), replicas are heated to different temperatures, while in REST2, the potential energy of a selected region (often the solute) is scaled to create effectively "hotter" replicas without physically heating the solvent [1] [40]. These methods rely on periodic exchange attempts between neighboring replicas, accepted with a probability that preserves detailed balance:
[ P_{\text{accept}} = \min\left(1, \exp\left(-\Delta\right)\right) ]
where (\Delta) depends on the potential energy difference in T-REMD or the scaled Hamiltonian difference in REST2. Efficient sampling requires adequate overlap of potential energy distributions between adjacent replicas, enabling frequent exchanges and ensuring all replicas perform a random walk through parameter space [39].
The Replica Segregation Problem
Replica segregation occurs when exchanges between adjacent replicas fail repeatedly, causing each replica to remain confined to its original parameter set. This breakdown manifests as poor "replica round-trip time"—the duration for a replica to traverse from one end of the parameter ladder to the other and back [39]. The consequences include:
- Inefficient sampling: Replicas become trapped in local minima corresponding to their specific parameters
- Poor convergence: Individual replicas fail to adequately explore conformational space
- Resource waste: Computational resources are expended without thermodynamic benefit
Theoretical analyses indicate that segregation risk increases with system size and complexity, larger parameter gaps between replicas, and insufficient simulation time to achieve proper equilibration [39] [40].
Comparative Performance Analysis
Exchange Efficiency and Sampling Performance
Table 1: Exchange Efficiency Metrics Across Enhanced Sampling Methods
| Method | System Type | Replica Count | Acceptance Rate | Round-Trip Time | Key Advantage |
|---|---|---|---|---|---|
| T-REMD | Small proteins/peptides | Scales with √(N atoms) | ~25% (achievable) | Highly system-dependent | Well-established methodology |
| REST2 | Biomolecular solutes | Reduced vs. T-REMD | Variable (15-30%) | Faster than T-REMD for focused regions | Targeted solute enhancement |
| ACES | Protein-ligand complexes | Similar to REST2 | Improved overlap via counter-diffusion | Optimized via dual topology | Balanced environmental response |
The fundamental trade-off in replica exchange methods lies between system size and computational feasibility. T-REMD requires replicas scaling with the square root of the system's degrees of freedom, becoming prohibitively expensive for large solvated systems [1]. REST2 addresses this by focusing enhancement on the solute, significantly reducing the required replica count while maintaining effective sampling of biologically relevant regions [40].
Experimental data from protein-ligand systems demonstrates that conventional REST2 implementations achieve moderate exchange rates (15-30%) but can suffer from environmental distortion—where the "hot" solute induces unnatural rearrangements in the surrounding protein or solvent [40]. This environmental response differential directly contributes to replica segregation by reducing phase space overlap between adjacent replicas.
Performance in Challenging Systems
Table 2: Method Performance in Specific Biological Contexts
| Method | Application | Sampling Acceleration | Key Limitation | Experimental Validation |
|---|---|---|---|---|
| REST2 | Aβ aggregation inhibition [41] | Enables μs-ms processes in ns-μs simulation | Potential force field inaccuracies | MM/PBSA binding free energies |
| REST2 | Mini-proteins (CLN025) [3] | Comparable to T-REMD with fewer replicas | Undersampling of high barriers | Free energy surface reconstruction |
| ACES | T4-lysozyme & Cdk2 ligands [40] | Superior to REST2 for rotamer sampling | Complex parameter optimization | Experimental crystal structure comparison |
| True RC Biasing | HIV-1 protease [14] | 10¹⁵-fold acceleration for flap opening | Requires reaction coordinate identification | Natural transition pathway reproduction |
Recent benchmarks reveal that while REST2 successfully captures conformational transitions in systems like Aβ trimers, enabling the study of amentoflavone's inhibitory mechanism [41], it struggles with high free-energy barriers without additional enhancements [3]. The integration of REST2 with diffusion-based generative models demonstrates improved performance in mapping conformational free-energy landscapes of enzymes like PTP1B, uncovering loop transition pathways consistent with biased simulations [3].
The ACES (Alchemically Enhanced Sampling) method explicitly addresses replica segregation by implementing a dual-topology framework that creates counter-balancing replica exchange networks, effectively minimizing environmental response differentials [40]. In direct comparisons, ACES demonstrates superior robustness to REST2 in handling conformational transitions in T4-lysozyme and Cdk2 ligand rotamer states where traditional MD and REST2-like methods fail [40].
Experimental Protocols and Methodologies
Standard REST2 Implementation Protocol
System Preparation:
- Force Field Selection: Employ biomolecule-appropriate force fields (CHARMM36m for proteins [41], AMBER for general biomolecules)
- Solvation: Utilize explicit solvent models (TIP3P water) with ion concentration matching physiological conditions
- Replica Parameterization: Define Hamiltonian scaling factors for solute regions using established protocols [40]
Simulation Parameters:
- Replica Count: Typically 24-40 replicas based on system size [41]
- Temperature Distribution: Logarithemically spaced between 310-389 K for biological systems [41]
- Exchange Attempt Frequency: 2.0 ps intervals to balance communication overhead and exchange probability [41]
- Simulation Duration: 100-200 ns per replica after equilibration [41]
Analysis Framework:
- Exchange Probability: Monitor acceptance rates between adjacent replicas
- Round-Trip Time: Track replica movement through parameter space
- Convergence Metrics: Assess via potential energy distributions and structural observables
Specialized Implementations Addressing Segregation
DDPM-Augmented REST2 [3]: This hybrid approach combines REST2 with Denoising Diffusion Probabilistic Models (DDPMs) to enhance sampling of high-barrier regions. The methodology involves:
- Running conventional REST2 simulations to obtain initial conformational ensembles
- Training DDPMs on the joint probability distribution in configuration and rescaled potential energy space
- Generating new configurations with accurate Boltzmann weights
- Iteratively refining free-energy surfaces through importance sampling
ACES Method [40]: The Alchemically Enhanced Sampling protocol specifically counters replica segregation through:
- Creation of enhanced sampling states by selectively turning off potential energy terms
- Implementation of dual topology framework with counter-balancing HREMD networks
- Application of smoothstep softcore potentials and non-linear Hamiltonian mixing
- Utilization of optimized λ-scheduling for improved phase space overlap
Diagram: Methodological Approaches to Replica Segregation
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Tools for Enhanced Sampling Studies
| Tool Category | Specific Solution | Function | Implementation Considerations |
|---|---|---|---|
| Simulation Software | GROMACS [41], AMBER [40], GENESIS [1] | MD engine with enhanced sampling capabilities | GPU acceleration critical for throughput |
| Enhanced Sampling Methods | REST2 [1], ACES [40], gREST [1] | Accelerate conformational transitions | Method selection depends on system size |
| Free Energy Analysis | MM/PBSA [41], MBAR [3], DDPM [3] | Estimate binding affinities and landscapes | DDPMs show promise for barrier regions |
| Force Fields | CHARMM36m [41], AMBER [40], CGenFF [41] | Define molecular interactions | IDP-optimized versions available |
| Analysis Tools | MDTraj, PyEMMA, GROMACS built-ins [41] | Process trajectories and quantify states | Automated pipelines improve reproducibility |
The replica segregation problem represents a fundamental challenge in enhanced sampling methodologies, directly impacting the efficiency and reliability of conformational sampling in drug discovery research. While REST2 provides significant advantages over temperature-based replica exchange through its targeted approach, it remains susceptible to exchange failures when environmental response differentials disrupt replica overlap.
Comparative analysis demonstrates that emerging methods like ACES and DDPM-augmented REST2 offer promising solutions to the segregation problem. ACES addresses the root cause through sophisticated alchemical pathways and dual-topology counter-diffusion networks [40], while DDPM integration enhances sampling of high-barrier regions that remain challenging for conventional REST2 [3]. For researchers selecting enhanced sampling strategies, the optimal approach depends critically on system characteristics: REST2 provides general-purpose enhancement for biomolecular solutes, ACES excels in protein-ligand systems with complex rotamer landscapes, and DDPM-augmented methods show promise for mapping elusive transition pathways.
Future methodological development will likely focus on adaptive parameter optimization, deeper integration of generative models, and improved force field accuracy—particularly for challenging systems like intrinsically disordered proteins where sampling limitations remain most pronounced. Through continued addressing of the replica segregation problem, enhanced sampling methodologies will expand their utility in illuminating protein function and accelerating therapeutic development.
Table of Contents
- Introduction: The Challenge of Biomolecular Sampling
- The Evolution of REST Protocols
- Methodological Comparison: REST2 vs. REST3
- Performance Analysis: Efficiency and Conformational Sampling
- Experimental Protocols for Key Studies
- Research Reagent Solutions
- Conclusions and Future Directions
Atomistic simulations of proteins in explicit solvent are a cornerstone of modern computational biology, yet capturing their large-scale conformational fluctuations remains a formidable challenge due to high energy barriers that lead to kinetic trapping and quasi-ergodicity in standard molecular dynamics (MD) simulations [9] [2]. Enhanced sampling techniques are therefore critical, particularly for studying intrinsically disordered proteins (IDPs) that exist as heterogeneous structural ensembles and rely on conformational plasticity for their function [9] [42]. Temperature Replica Exchange (T-RE) is a powerful method that facilitates barrier crossing by allowing replicas of the system to perform a random walk in temperature space [9]. However, its application to explicit solvent simulations is hampered by poor scaling with system size; the number of replicas required grows with the square root of the total number of atoms, making simulations of even modestly sized solvated systems computationally prohibitive [9] [16].
Replica Exchange with Solute Tempering (REST) was developed to overcome this limitation. The core idea is to apply effective tempering only to a selected "solute" region, thereby drastically reducing the number of degrees of freedom that contribute to the replica exchange acceptance probability [9] [16]. While the original REST (retroactively termed REST1) and its improved version, REST2, demonstrated significant speedups, studies revealed that REST2 promotes an artificial conformational collapse in intrinsically disordered proteins (IDPs) at high effective temperatures [9]. This collapse creates an exchange bottleneck, hindering sampling. This discovery motivated the recent development of REST3, which re-calibrates solute-solvent interactions to correct this bias and enable more efficient exploration of biomolecular conformational landscapes [9] [43].
The Evolution of REST Protocols
The REST methodology aims to enhance the sampling of a solute, such as a protein, while simulating the entire system (solute and explicit solvent) at a single physical temperature. This is achieved by scaling different components of the potential energy function across replicas.
REST2: This protocol scales the solute-solute (
Epp) and solute-solvent (Epw) interaction energies by a factor ofβ_m/β_0(whereβ_m = 1/kBT_m), while the solvent-solvent (Eww) interactions remain unscaled [9] [16]. The scaling of theEpwterm intentionally weakens the protein-water interactions at higher effective temperatures. This was designed to maintain compact protein conformations to facilitate the refolding of small, structured proteins and peptides [16].REST3: The REST3 protocol introduces a crucial modification to address a key limitation of REST2. It incorporates an additional calibration factor,
κ_m, specifically for the van der Waals (vdW) component of the solute-solvent interactions [9] [43]. The potential energy for a replica m in REST3 is given by:E_m^REST3(X) = (β_m/β_0)E_pp(X) + (β_m/β_0)E_pw^elec(X) + κ_m(β_m/β_0)E_pw^vdW(X) + E_ww(X)This calibrated vdW scaling counteracts the excessive weakening of solute-solvent interactions in REST2, preventing the artificial collapse of IDPs at high effective temperatures and promoting a more realistic chain expansion [9].
The following diagram illustrates the logical progression and core differences between these REST variants:
Methodological Comparison: REST2 vs. REST3
The core difference between REST2 and REST3 lies in the treatment of solute-solvent van der Waals interactions, which directly impacts the conformational ensemble of the solute.
Table 1: Hamiltonian Scaling in REST2 and REST3 Protocols
| Energy Component | REST2 Scaling Factor | REST3 Scaling Factor |
|---|---|---|
Solute-Solute (Epp) |
β_m/β_0 |
β_m/β_0 |
Solute-Solvent Electrostatics (Epw^elec) |
β_m/β_0 |
β_m/β_0 |
Solute-Solvent vdW (Epw^vdW) |
β_m/β_0 |
κ_m * (β_m/β_0) |
Solvent-Solvent (Eww) |
1 (unscaled) |
1 (unscaled) |
Table 2: Conformational and Sampling Implications
| Feature | REST2 | REST3 |
|---|---|---|
| Solute-Solvent Interactions | Weakened at high effective temperatures | Re-calibrated vdW to maintain realistic interactions |
| IDP Conformations at High T | Artificially compact and collapsed | Realistic level of chain expansion |
| Replica Exchange | Can lead to segregation and poor random walk | More efficient temperature random walk |
| Primary Application | Reversible folding of small, structured proteins | Sampling of disordered and flexible proteins |
Performance Analysis: Efficiency and Conformational Sampling
Empirical studies on intrinsically disordered proteins (IDPs) like the p53 N-terminal domain (p53-NTD) and the CREB transactivation domain have quantitatively demonstrated the advantages of REST3.
Eliminating Artificial Collapse: REST2 was found to promote overly compact conformations of IDPs at high effective temperatures, causing replicas to become segregated and hindering the random walk necessary for effective sampling [9]. REST3's parameter
κ_mwas specifically tuned to reproduce realistic levels of protein chain expansion, thereby eliminating this exchange bottleneck [9].Improved Sampling Efficiency: In direct comparisons, REST3 leads to a much more efficient temperature random walk than REST2. This enhanced replica mobility translates to improved convergence of conformational ensembles [9]. The increased efficiency is so significant that REST3 can achieve similar or better conformational convergence than REST2 using a smaller number of replicas, offering direct computational savings [9].
Quantitative Benchmarking: The performance gain is quantifiable through metrics like replica exchange acceptance rates and the rate of diffusion of replicas through temperature space. REST3 consistently shows superior performance in these metrics for systems involving large-scale conformational fluctuations [9].
The typical workflow for a comparative REST2/REST3 study, from system setup to analysis, is outlined below:
Experimental Protocols for Key Studies
The experimental data cited in this guide primarily stems from studies evaluating REST2 and REST3 on intrinsically disordered proteins. Below is a summary of a typical computational methodology.
Table 3: Representative Experimental Protocol for REST2/REST3 Comparison
| Protocol Step | Description |
|---|---|
| System Preparation | Proteins: Intrinsically disordered proteins (IDPs) such as the p53 N-terminal domain (residues 1-61) or the kinase inducible transactivation domain of CREB [9]. Solvation: Explicit solvent (e.g., TIP3P water model) in a simulation box with sufficient padding to accommodate extended conformations [9]. System Size: ~72,000 atoms for p53-NTD [9]. |
| Replica Parameters | Number of Replicas: ~16 replicas for a temperature range of 298 K to 500 K [9]. REST3 may achieve similar acceptance rates with fewer replicas [9]. Effective Temperature Spacing: Exponentially spaced between the physical temperature (T0) and a maximum effective temperature (Tmax), calculated as T_m = T_0 (T_max/T_0)^(m/(M-1)) for replica m out of M total replicas [9]. |
| Simulation Details | Software: MD packages with REST capability (e.g., GROMACS, AMBER, NAMD, OpenMM). Force Fields: Modern force fields balanced for disordered proteins (e.g., AMBER ff99SB-ILDN, CHARMM36m) [9] [42]. Simulation Length: Multi-nanosecond production runs per replica after equilibration. Exchange attempts typically every 1-2 ps [9]. |
| Analysis Metrics | Sampling Efficiency: Replica exchange acceptance rates and round-trip time in temperature space [9]. Conformational Properties: Radius of gyration (Rg), end-to-end distance, and secondary structure propensity [9]. Convergence is assessed by the overlap between independent estimates from multiple simulations [2]. |
Research Reagent Solutions
This section details key software, force fields, and models essential for implementing and running REST simulations.
Table 4: Essential Research Reagents for REST Simulations
| Reagent / Solution | Function / Description | Examples / Notes |
|---|---|---|
| MD Simulation Software | Engine for running molecular dynamics and replica exchange. | GROMACS [16], AMBER, NAMD, OpenMM. Must support Hamiltonian replica exchange and the desired REST variant. |
| Force Fields | Mathematical functions and parameters defining interatomic interactions. | AMBER [44], CHARMM [16], OPLS [14], GROMOS [3]. Select versions rebalanced for IDPs (e.g., CHARMM36m, AMBER ff99SB-ILDN) [9] [42]. |
| Water Models | Represent explicit solvent molecules. | TIP3P [9], SPC/E, TIP4P. TIP3P is commonly used in biomolecular simulations. |
| Analysis Tools | Software for processing simulation trajectories and calculating metrics. | MDTraj, PyEMMA, CPPTRAJ, GROMACS analysis suite, VMD [2]. Used for calculating Rg, RMSD, and free energies. |
| Enhanced Sampling Plugins | Libraries that provide advanced sampling algorithms. | PLUMED [43] [14] is a widely used plugin for adding biasing potentials and analyzing collective variables. |
The development from REST2 to REST3 highlights a critical principle in enhanced sampling: the precise balance of solute-solvent interactions is paramount for generating realistic conformational ensembles, especially for flexible biomolecules like IDPs. While REST2 remains a powerful tool for studying the folding of small, structured proteins, REST3 emerges as a superior protocol for sampling the heterogeneous landscapes of intrinsically disordered proteins and large-scale conformational changes by preventing artificial collapse and enabling more efficient replica exchange [9].
Looking forward, the integration of REST with other advanced sampling and analysis techniques represents the cutting edge of the field. Future developments are likely to focus on several key areas, as visualized below:
These hybrid approaches, which combine the broad Hamiltonian scaling of REST with targeted biasing or machine-learning-driven analysis, promise to further overcome the entropic barriers that limit pure tempering methods, unlocking access to increasingly complex biomolecular processes [9] [45] [14].
Integrating REST2 with Free Energy Perturbation (FEP) and Umbrella Sampling
In molecular dynamics (MD) simulations, overcoming kinetic traps and achieving sufficient conformational sampling remains a significant challenge for calculating free energies in complex biomolecular systems. Enhanced sampling techniques are essential for obtaining statistically robust results in practical computation times. This guide focuses on the integration of the Replica Exchange with Solute Tempering 2 (REST2) method with two foundational free energy methods: Free Energy Perturbation (FEP) and Umbrella Sampling (US). Within the broader thesis of comparing enhanced sampling approaches, REST2 offers a specific advantage through its Hamiltonian scaling approach, which selectively accelerates the sampling of a designated "solute" region, leading to more efficient exploration of phase space compared to standard temperature-based replica exchange or conventional MD simulations. We will objectively compare the performance of REST2-enhanced protocols against alternative methods, providing structured experimental data and implementation details to inform researchers and drug development professionals.
Theoretical Foundation and Methodology
The REST2 Framework
Replica Exchange with Solute Tempering 2 (REST2) is an enhanced sampling method that belongs to the class of Hamiltonian Replica Exchange (H-REM) techniques. Its core innovation lies in scaling the potential energy terms associated with a specific "solute" or "hot region" across different replicas, while all replicas are simulated at the same physical temperature [16]. This contrasts with Temperature Replica Exchange (T-REM), where the entire system's temperature is varied, leading to a rapid increase in the required number of replicas with system size.
In REST2, the potential energy for a given replica ( m ) is defined as [16]: [ Em^{REST2}(X) = \frac{\betam}{\beta0}E{pp}(X) + \sqrt{\frac{\betam}{\beta0}}E{pw}(X) + E{ww}(X) ] Here, ( X ) represents the system configuration, ( E{pp} ) is the solute-solute interaction energy, ( E{pw} ) is the solute-solvent interaction energy, and ( E{ww} ) is the solvent-solvent interaction energy. The factors ( \betam = 1/kB Tm ) and ( \beta0 = 1/kB T0 ), where ( T0 ) is the target temperature of interest. This scaling effectively lowers energy barriers for the solute, facilitating faster conformational transitions. The acceptance probability for exchanging configurations between replicas ( m ) and ( n ) depends on the energy difference [16]: [ \Delta{mn}^{(REST2)} = (\betam - \betan)\left[(E{pp}(Xn) - E{pp}(Xm)) + \frac{\beta0}{\betam + \betan}(E{pw}(Xn) - E{pw}(Xm))\right] ] This formulation removes the dependence on the solvent-solvent interactions, allowing for a reduced number of replicas compared to T-REM.
Integration with Free Energy Calculation Methods
REST2 with Free Energy Perturbation (FEP)
Free Energy Perturbation is an alchemical method for calculating free energy differences by transforming one system into another along a non-physical pathway. Integrating REST2 with FEP involves using the replica exchange framework to enhance the sampling of the alchemical intermediate states [19]. The "hot region" in this context typically includes the perturbed atoms of the ligand and often key protein residues in the binding site. The Hamiltonian for each replica includes both the alchemical parameter ( \lambda ) and the REST2 scaling, creating a 2D replica exchange lattice that facilitates sampling in both conformational and alchemical space. This combination helps overcome sampling barriers that plague standard FEP simulations, such as rotameric transitions of ligands or side chains.
REST2 with Umbrella Sampling (US)
Umbrella Sampling is a biased sampling technique used to calculate free energy landscapes along pre-defined collective variables (CVs). It involves running multiple simulations (windows), each with a restraining potential that forces the system to sample a specific region of the CV space. Integrating REST2 with US involves running a REST2 simulation within each umbrella window [19]. The "hot region" is chosen to include the degrees of freedom most relevant to the CV. This hybrid approach, sometimes called US/REST2, enhances the conformational sampling within each window, ensuring better convergence of the free energy profile, especially for complex transitions involving large biomolecular rearrangements.
Performance Comparison and Experimental Data
The following tables summarize key performance metrics from various studies comparing REST2-integrated methods against standard and alternative enhanced sampling techniques.
Table 1: Performance Comparison for Peptide Folding Simulations
| Method | System | Number of Replicas | Sampling Efficiency (Relative to T-REM) | Key Observation | Citation |
|---|---|---|---|---|---|
| T-REM | Trpcage, β-hairpin | Scales with ( \sqrt{f} ) (Total DOF) | 1.0 (Baseline) | Folding achievable but computationally expensive | [16] |
| REST1 | Trpcage, β-hairpin | Scales with ( \sqrt{f_p} ) (Solute DOF) | Lower than T-REM | Poor exchange between folded/unfolded states | [16] |
| REST2 | Trpcage, β-hairpin | Scales with ( \sqrt{f_p} ) (Solute DOF) | Higher than REST1 & T-REM | Efficient folding/unfolding transitions; robust sampling | [16] |
| ACES | Cdk2 Ligand, T4L L111 | Not Specified | Superior to REST2 | Handled different rotamer states and side-chain distributions | [40] |
Table 2: Performance in Free Energy Calculations and Protein-Ligand Systems
| Method | Application | Performance & Outcome | Comparison to Alternative Methods | Citation |
|---|---|---|---|---|
| FEP/REST2 | Absolute Binding Affinity (Protein-Ligand) | Quantitative binding affinity calculation | Enabled sampling of relevant configurations not efficiently sampled by standard FEP | [19] |
| US/REST2 | Free Energy Landscape Exploration | Improved convergence of free energy profiles | Enhanced sampling within each umbrella window compared to standard US | [19] |
| ACES | Hydration Free Energy (Acetic Acid) | Result independent of starting conformation | Superior to traditional MD and REST2-like methods in robustness | [40] |
| ACES | Hydration Free Energy (FreeSolv molecules) | Closer agreement with experiment | Corrected outliers from standard database calculations | [40] |
| REST2 | Peptide Conformation (Ac-(AAQAA)3-NH2) | Sampled folded/unfolded states in explicit water | Achieved with 16 replicas (300-600 K range); demonstrated practical utility | [19] |
Key Insights from Comparative Data:
- Efficiency: REST2 significantly reduces the number of replicas required compared to T-REM by scaling with the square root of the solute's degrees of freedom rather than the entire system's [16] [19].
- Superior Sampling: For systems with large conformational changes (e.g., peptide folding), REST2 demonstrates higher sampling efficiency and more robust exploration of phase space than its predecessor, REST1 [16].
- Robustness in Free Energy Calculations: Integrated with FEP and US, REST2 helps achieve more convergent and reliable free energy estimates by overcoming kinetic traps, as shown in protein-ligand binding and hydration free energy studies [40] [19].
- Emerging Alternatives: The recently developed Alchemically Enhanced Sampling (ACES) method has shown superior performance to REST2 in specific test cases, particularly in handling rotameric states and protein side-chain distributions, indicating a vibrant field of ongoing development [40].
Experimental Protocols
Protocol: FEP/REST2 for Absolute Binding Affinity
This protocol outlines the key steps for performing an absolute binding free energy calculation using FEP augmented with REST2, as implemented in NAMD [19].
System Setup:
- Prepare the protein-ligand complex in its explicit solvent environment (e.g., TIP3P water box with ions).
- Generate the topology and parameter files for the ligand, ensuring compatibility with the chosen force field.
Define the Alchemical Pathway:
- Create a set of ( \lambda ) states that gradually annihilate (or decouple) the ligand from its environment in the binding site. A typical number of ( \lambda ) windows ranges from 12 to 20.
- Employ soft-core potentials to avoid singularities as atoms are annihilated.
Define the REST2 "Hot Region":
- Select the atoms to be scaled by REST2. This region should include the entire ligand and potentially key protein residues in the binding site that may undergo conformational changes.
- Using visualization software like VMD, create a selection and output it to a PDB file to be read by the simulation software.
Set Up the Replica Exchange Lattice:
- Establish a set of replicas that vary in both the alchemical ( \lambda ) parameter and the REST2 scaling factor ( \betam / \beta0 ). This creates a 2D grid for replica exchange.
- The number of REST2 replicas is determined by the desired temperature range for the "hot region" (e.g., from the target temperature ( T0 ) to a maximum ( T{max} )) and the targeted exchange acceptance rate.
Simulation Execution:
- Run the multi-replica simulation on a high-performance computing (HPC) platform. Each replica is assigned to a separate group of CPU/GPU cores.
- Configure the simulation for periodic exchange attempts between neighboring replicas in both the alchemical and REST2 dimensions. A high attempt frequency (e.g., every 1-2 ps) is recommended for optimal sampling [19].
Analysis:
- Use the Multistate Bennett Acceptance Ratio (MBAR) or the Weighted Histogram Analysis Method (WHAM) on the collected data from all replicas to compute the final absolute binding free energy.
- Assess convergence by monitoring the time evolution of the free energy estimate and the replica exchange statistics.
Protocol: US/REST2 for Free Energy Landscape
This protocol describes how to combine Umbrella Sampling with REST2 to calculate a free energy profile along a collective variable [19].
Collective Variable (CV) Selection:
- Define a CV that accurately describes the process of interest (e.g., a distance, angle, or root-mean-square deviation).
Umbrella Sampling Windows:
- Perform a preliminary simulation (e.g., steered MD) to generate initial configurations along the CV.
- Define a set of umbrella windows that collectively cover the entire range of the CV. Each window has a harmonic restraint centered at a specific value of the CV.
Integrate REST2:
- For each umbrella window, set up a separate REST2 simulation. The "hot region" should be selected to include the molecular groups whose conformational changes are critical for the transition described by the CV.
Run REST2-Enhanced Umbrella Simulations:
- Execute the simulations for all umbrella windows concurrently. Within each window, multiple REST2 replicas (with different scaling factors) run and periodically attempt exchanges.
- This setup enhances the sampling of side-chain motions, rotameric states, and other local conformational changes within the geographic constraint of the umbrella potential.
Free Energy Reconstruction:
- After the simulations, use WHAM or MBAR to unbias the restraints from all umbrella windows and all REST2 replicas, combining the data to produce a one-dimensional or two-dimensional potential of mean force (PMF).
- The improved sampling within windows often leads to a faster converging and more reliable PMF compared to standard US.
Workflow Visualization
US/REST2 Workflow: This diagram outlines the sequential workflow for combining Umbrella Sampling with the REST2 enhanced sampling method.
REST2 Scaling Logic: This diagram visualizes the core REST2 concept of creating multiple scaled replicas of the solute Hamiltonian to enhance conformational sampling.
Comparative Enhanced Sampling Framework: This diagram places REST2 within a hierarchy of enhanced sampling methods, highlighting its conceptual advances and relationship to alternatives like ACES.
Table 3: Key Software and Computational Resources for REST2 Simulations
| Tool/Resource | Category | Primary Function/Description | Relevant Citation |
|---|---|---|---|
| NAMD | MD Simulation Software | Highly scalable MD program; a common platform for implementing REST2 and running FEP/US simulations. | [19] |
| AMBER | MD Simulation Software | Suite of biomolecular simulation programs; includes tools for alchemical free energy calculations (e.g., ACES method). | [40] |
| VMD | Visualization & Analysis | Used to prepare simulation systems, select atoms for the REST2 "hot region," and visualize trajectories. | [19] |
| Charm++ | Parallel Programming System | The underlying architecture of NAMD that enables efficient parallel communication and replica exchange. | [19] |
| MBAR/WHAM | Analysis Tool | Statistical methods for analyzing data from multi-state simulations (e.g., FEP, US) to compute free energies. | [19] |
| IBM Blue Gene/Q | HPC Infrastructure | Example of a high-performance computing system used for large-scale REST2 simulations. | [19] |
Benchmarking Performance: How REST2 Stacks Up Against Standard MD and Other Methods
Molecular dynamics (MD) simulations are a cornerstone of modern computational biology, providing atomic-level insights into biomolecular processes. However, their utility is often limited by the computational expense required to achieve sufficient conformational sampling. Enhanced sampling techniques like the Temperature Replica Exchange Method (TREM) address this by accelerating barrier crossing, but their computational cost scales poorly with system size. Replica Exchange with Solute Tempering 2 (REST2) has emerged as a powerful alternative that significantly reduces computational demands compared to standard TREM [16] [19]. This guide provides a objective performance comparison between REST2 and standard MD conformational sampling methods, quantifying the reductions in CPU time and replica count, which are critical metrics for research efficiency in academic and industrial drug development.
Understanding the Methods and Their Evolution
Standard Temperature Replica Exchange (TREM)
In TREM, multiple non-interacting copies (replicas) of the system are simulated simultaneously at different temperatures. Periodically, exchanges between neighboring temperatures are attempted and accepted according to the Metropolis criterion, which ensures detailed balance. This allows a replica to perform a random walk in temperature space, effectively overcoming kinetic traps. However, a significant limitation is that the number of replicas required for a given acceptance ratio scales with the square root of the system's degrees of freedom (f). For a solvated protein system, this translates to a requirement for dozens or even hundreds of replicas, making simulations of large biomolecules computationally prohibitive [16] [45].
Replica Exchange with Solute Scaling (REST2)
REST2 is a Hamiltonian Replica Exchange (H-REX) method designed to overcome TREM's poor scaling. Instead of heating the entire system (solute and solvent), REST2 selectively scales the Hamiltonian of the solute and its interactions with the solvent. All replicas are run at the same physical temperature, but the potential energy function for replica m is scaled as follows [16] [19]:
E_m^REST2(X) = (β_m / β_0) * E_pp(X) + (β_m / β_0) * E_pw(X) + E_ww(X)
Here, E_pp, E_pw, and E_ww represent solute-solute, solute-solvent, and solvent-solvent interaction energies, respectively. β_m = 1/k_B T_m and β_0 = 1/k_B T_0, where T_0 is the target temperature and T_m is an effective "temperature" for the solute. This scaling lowers energy barriers for the solute in higher replicas, enhancing its conformational sampling. Crucially, because solvent-solvent interactions (E_ww) are identical in all replicas and do not contribute to the exchange acceptance probability, the number of required replicas scales with the square root of the solute's degrees of freedom (f_p), not the entire system's [16].
Table: Key Conceptual Differences Between TREM and REST2
| Feature | Temperature Replica Exchange (TREM) | Replica Exchange with Solute Scaling (REST2) |
|---|---|---|
| Scaling Principle | Temperature of the entire system is varied. | Hamiltonian of the solute and solute-solvent interactions is scaled; all replicas at same physical temperature. |
| Replica Count Scaling | Scales as √f, where f is the total system degrees of freedom. |
Scales as √f_p, where f_p is the solute degrees of freedom. |
| Computational Focus | Enhances sampling of all system degrees of freedom. | Selectively enhances sampling of the solute's conformational space. |
| Solvent Treatment | Solvent is "hot" in high-temperature replicas. | Solvent remains "cold" in all replicas. |
The Scientist's Toolkit: Essential Components for REST2
Table: Key Research Reagents and Software for REST2 Simulations
| Item | Function in REST2 Simulation |
|---|---|
| MD Engine with REST2 | Software like NAMD provides the computational framework to run the MD simulations, handle the REST2 Hamiltonian scaling, and manage replica exchanges [19]. |
| System Builder | Tools like VMD are used to prepare the initial molecular system, including solvation and ionization, and to define the "hot" solute region for REST2 [19]. |
| Force Field | A set of potential functions (e.g., CHARMM, AMBER) defining interatomic forces. REST2 scales specific force field parameters (charges, LJ ε) of the solute atoms [19]. |
| Parallel Computing Cluster | High-performance computing infrastructure is essential to run multiple replicas concurrently, enabling the replica exchange process. |
Quantitative Performance Comparison
Experimental data from peer-reviewed studies consistently demonstrates REST2's superior computational efficiency across various biomolecular systems.
Direct Comparisons of Replica Count and Sampling
A foundational study by Wang et al. compared REST2, TREM, and the original REST (REST1) for folding the Trp-cage and β-hairpin peptides in explicit water. The results confirmed that REST2 "greatly reduces the number of CPUs required by regular replica exchange" and "greatly increases the sampling efficiency over REST1" [16] [46]. This is attributed to the more effective lowering of solute energy barriers in REST2 and a more favorable acceptance probability formula that facilitates better replica mixing [16].
Further benchmarking on the CLN025 mini-protein showed that a method combining REST2 with diffusion models "achieved comparable accuracy to TREM while requiring fewer replicas" [17]. This highlights that the efficiency gains of REST2 do not necessarily come at the cost of accuracy.
Table: Experimental Reductions in Replica Count and CPU Time
| System Studied | Standard TREM Replicas | REST2 Replicas | Reduction & Efficiency Gain |
|---|---|---|---|
| Trp-cage & β-hairpin folding [16] | Required a large number (N/A) as per √f scaling. |
Required a much smaller number (N/A). | Greatly reduced CPU count and higher sampling efficiency vs. TREM and REST1. |
| p53 N-terminal domain (IDP) [9] | Estimated >100 replicas for ~20% acceptance. | 16 replicas used with ~25% acceptance. | ~6-fold reduction in replicas (from ~100+ to 16) for a system of ~72,000 atoms. |
| CLN025 mini-protein [17] | N/A (Used as a benchmark). | Achieved comparable accuracy with fewer replicas. | Fewer replicas required to achieve accuracy comparable to TREM. |
| Ac-(AAQAA)₃-NH₂ peptide [19] | N/A (System with ~25,000 atoms). | 16 replicas used spanning 300–600 K effective temp. | Demonstrated practical application with a feasible number of replicas for a solvated system. |
Protocol for Key Efficiency Experiments
The quantitative data in the table above are derived from rigorous simulation protocols. A representative methodology for a REST2 efficiency study is outlined below [16] [19]:
- System Preparation: The protein (e.g., Trp-cage, β-hairpin, Ac-(AAQAA)₃-NH₂) is placed in an explicit water box, and the system is neutralized with ions.
- Replica Setup:
- For TREM, a series of temperatures is chosen, typically exponentially spaced between a low (e.g., 300 K) and a high temperature (e.g., 500-600 K). The number of replicas is determined to ensure a sufficient exchange acceptance rate (e.g., >20%) over the entire temperature range.
- For REST2, a series of effective solute temperatures (
T_m) is chosen, also exponentially spaced betweenT_0(e.g., 300 K) andT_max(e.g., 600 K). The number of replicas is based on the solute's degrees of freedom, which is significantly lower.
- Simulation Parameters: All replicas are minimized and equilibrated. Production simulations are run using a MD engine like NAMD. In REST2, the "hot region" is defined, and its force field parameters (charges, Lennard-Jones ε) are scaled on-the-fly by a factor of
√(β_m/β_0)and(β_m/β_0), respectively [19]. - Exchange Attempts: Exchanges between neighboring replicas are attempted periodically (e.g., every 1-2 ps). The acceptance probability is calculated using the Metropolis criterion based on the specific energy differences for each method (e.g., Eq. 4 from [16] for REST2).
- Efficiency Metrics:
- Replica Count: The minimum number of replicas required to maintain a target exchange acceptance rate (e.g., 20-30%) across the entire temperature/Hamiltonian ladder is recorded.
- Sampling Efficiency: This is assessed by measuring the rate of folding/unfolding transitions, the convergence of structural properties (e.g., RMSD, radius of gyration) over time, and the quality of the resulting free energy landscapes compared to experimental data or benchmark simulations.
The experimental data conclusively demonstrates that REST2 provides a substantial efficiency advantage over standard TREM for conformational sampling of biomolecules in explicit solvent. The core of this advantage lies in the drastic reduction in the number of replicas required, which translates directly into lower computational cost and CPU time. For researchers and drug development professionals, this efficiency gain means that more complex systems, such as protein-ligand complexes or large intrinsically disordered proteins, can be studied with enhanced sampling techniques at a feasible computational cost, accelerating the pace of scientific discovery and in silico drug design.
In computational biochemistry, achieving robust thermodynamic averages is the cornerstone for obtaining reliable insights into biomolecular function, drug binding, and conformational dynamics. The core challenge lies in the ergodic hypothesis, which assumes that a simulation will sample all conformations accessible to the system with a probability proportional to their Boltzmann weight. In practice, the rugged, high-dimensional free energy landscapes of biomolecules feature numerous metastable states separated by kinetic barriers that are often insurmountable within the timescales of conventional Molecular Dynamics (MD) simulations. This leads to quasi-ergodicity, where the simulation becomes trapped in local energy minima, resulting in poorly converged and statistically unreliable thermodynamic averages. The severity of this sampling problem escalates with system size and complexity, particularly for large proteins or intrinsically disordered proteins (IDPs) that explore a vast conformational landscape.
This guide provides a comparative analysis of sampling methodologies, focusing on the efficiency of the Replica Exchange with Solute Tempering 2 (REST2) algorithm against standard MD sampling techniques. The objective is to equip researchers with the data and protocols necessary to select and implement the most appropriate method for ensuring convergence in their specific studies, thereby deriving thermodynamic averages that are both accurate and statistically meaningful.
Methodological Comparison: REST2 vs. Standard MD Sampling
Standard temperature-based sampling methods, like the Temperature Replica Exchange Method (TREM), enhance sampling by running multiple replicas of the system at different temperatures. High-temperature replicas can overcome energy barriers, and configuration exchanges between replicas facilitate a random walk through temperature space. However, a significant limitation is that the number of replicas required scales with the square root of the system's degrees of freedom (O(√f)). Since the total energy (f) is dominated by the solvent in explicit water simulations, TREM becomes computationally prohibitive for large, solvated biomolecular systems [16] [2].
REST2, a Hamiltonian replica exchange method, addresses this bottleneck by focusing the enhanced sampling on the solute. In REST2, all replicas run at the same temperature, but the Hamiltonian of the solute is scaled. This effectively lowers the energy barriers within the solute, promoting conformational transitions.
Table 1: Fundamental Comparison of Sampling Methodologies
| Feature | Standard MD | Temperature Replica Exchange (TREM) | REST2 |
|---|---|---|---|
| Sampling Principle | Newtonian dynamics on the original potential energy surface. | Multiple temperatures to overcome barriers; exchanges between replicas. | Scaled solute Hamiltonian at a single temperature; exchanges between replicas. |
| Computational Scaling | N/A (base cost per nanosecond). | Scales as O(√f), where f is the total degrees of freedom (solute + solvent). |
Scales as O(√fp), where fp is the solute's degrees of freedom. |
| Key Advantage | Simplicity; directly generates dynamics. | Effective at overcoming barriers for small systems. | Superior scalability for large, solvated systems; reduces required CPUs. |
| Primary Limitation | Prone to quasi-ergodicity; poor sampling of rare events. | Number of replicas becomes prohibitively high for large systems. | Parameterization of the scaled Hamiltonian is critical for performance. |
The key innovation of REST2 is its scaling strategy. The potential energy for a replica m is defined as:
E_m^REST2(X) = (β_m/β_0)E_pp(X) + √(β_m/β_0)E_pw(X) + E_ww(X)
where E_pp is the solute intra-molecular energy, E_pw is the solute-solvent interaction energy, E_ww is the solvent-solvent energy, and β_m = 1/k_B T_m [16]. This scaling reduces the number of required replicas, as the acceptance probability for exchange depends only on the solute's energy terms, not on the vast number of solvent degrees of freedom.
Performance Benchmarking and Quantitative Comparison
Empirical studies across various protein systems demonstrate REST2's superior performance in achieving convergence. Benchmarking against both standard TREM and its predecessor (REST1) reveals significant gains.
Table 2: Performance Benchmarking on Model Systems
| Protein System | Comparison Method | Key Performance Metric | Result for REST2 |
|---|---|---|---|
| trpcage & β-hairpin [16] | TREM & REST1 | CPU time for ab initio folding | Greatly reduced vs. TREM; Greatly increased sampling efficiency vs. REST1 |
| trpcage & β-hairpin [16] | TREM | Number of replicas (CPUs) required | Fewer replicas required due to better scaling with system size |
| CLN025 mini-protein [3] | TREM | Accuracy of free-energy surface | Achieved comparable accuracy to TREM |
| PTP1B enzyme loop [3] | Biased sampling methods | Discovery of high-barrier transition pathways | Uncovered complex pathway with minimal computational overhead |
For the trpcage and β-hairpin proteins, which undergo large-scale conformational changes, the original REST1 method was found to be less efficient than TREM. However, the modified scaling in REST2 proved "much more efficient than REST1 in sampling the conformational space of large systems undergoing large conformation changes" [16]. The improvement stems from a more effective lowering of intra-solute energy barriers and a favorable cancellation of energy terms in the replica exchange acceptance criterion [16].
Furthermore, modern hybrid approaches are pushing the boundaries of REST2. A recent framework combined REST2 with Denoising Diffusion Probabilistic Models (DDPMs), a deep learning tool. This integration refines the free-energy landscape obtained from REST2 simulations, improving the resolution of high-barrier regions and enabling the discovery of complex transition pathways, as demonstrated for the PTP1B enzyme, with minimal added computational cost [3].
Detailed Experimental Protocols
To ensure reproducibility and robust convergence analysis, below are detailed protocols for key experiments cited in this guide.
Protocol 1: REST2 Simulation for Protein Folding
This protocol is adapted from studies on the trpcage and β-hairpin proteins [16].
System Setup:
- Initial Structure: Obtain or generate an initial folded or extended structure of the target protein (e.g., trpcage, PDB ID 1L2Y).
- Solvation: Place the protein in a dodecahedral simulation box with a minimum distance of 1.2 nm between the protein and box edge.
- Water Model: Use explicit water models such as TIP3P.
- Neutralization: Add ions (e.g., Na⁺/Cl⁻) to neutralize the system and then bring to a physiological salt concentration (e.g., 150 mM NaCl).
Force Field and Parameterization:
- Apply a modern biomolecular force field (e.g., CHARMM36, AMBER).
- For REST2, the solute's bonded terms (dihedral angles), Lennard-Jones ε parameters, and atomic charges are scaled by factors of
(β_m/β_0),(β_m/β_0), and√(β_m/β_0), respectively, for each replicam[16].
Simulation Parameters:
- Software: Use MD packages that support Hamiltonian replica exchange (e.g., GROMACS, AMBER, GENESIS).
- Replica Parameters: Typically, 8-16 replicas are used for REST2. The effective temperature range should be chosen so that the highest "temperature" replica can efficiently unfold and sample extended conformations. Exchanges between neighboring replicas are attempted every 1-2 ps.
- Equilibration: Perform energy minimization followed by a short NVT and NPT equilibration with position restraints on protein heavy atoms.
- Production Run: Conduct a production REST2 simulation for a duration sufficient to observe multiple folding/unfolding events (hundreds of nanoseconds to microseconds per replica).
Protocol 2: Convergence Analysis via Markov State Models (MSMs)
This protocol is used in studies like the analysis of KRAS activation [47] to derive thermodynamics and kinetics from simulation data.
Data Generation:
- Perform multiple, independent REST2 or MD simulations starting from different conformations (e.g., folded, unfolded, crystal structures).
- Save molecular trajectories at a regular interval (e.g., every 10-100 ps).
Featurization:
- Extract features from the trajectories that describe the slow conformational changes. Common features include inter-atomic distances, torsion angles, and root-mean-square deviation (RMSD) of specific regions.
Dimensionality Reduction:
- Use methods like Time-lagged Independent Component Analysis (tICA) to project the high-dimensional features into a lower-dimensional space that captures the slowest dynamical processes.
Clustering and Model Building:
- Cluster the data in the reduced space into discrete conformational states (microstates).
- Build a Markov State Model (MSM) by counting transitions between these states at a lag time. Validate the model by checking the implied timescales and conducting Chapman-Kolmogorov tests.
Analysis of Convergence:
- Thermodynamics: Analyze the stationary distribution of the MSM to identify metastable states (macrostates) and their equilibrium populations.
- Kinetics: Calculate the transition times and pathways between metastable states.
- Convergence Check: Verify that the observed state populations and implied timescales remain stable when the model is built using different subsets of the data or increasing simulation time.
Diagram 1: MSM Workflow for Convergence
Advanced Integration: Combining REST2 with Generative AI
A cutting-edge advancement to improve convergence, particularly in high free-energy barrier regions, is the fusion of REST2 with generative artificial intelligence. The following workflow illustrates this hybrid approach [3]:
Diagram 2: AI-Enhanced REST2 Sampling
- Initial REST2 Simulation: A standard REST2 simulation is performed, providing an initial, albeit potentially imperfect, sampling of the conformational landscape.
- Train DDPM on REST2 Data: A Denoising Diffusion Probabilistic Model (DDPM) is trained on the configurations from all REST2 replicas. The DDPM learns the joint probability distribution between the system's configurations and their scaled potential energy.
- Generate New Configurations: The trained DDPM can generate new, thermodynamically consistent configurations, effectively filling in gaps in the undersampled regions of the initial simulation.
- Importance Sampling on Collective Variables (CVs): For systems where key reaction coordinates are known, the generated configurations can be used to perform targeted importance sampling along these CVs, further refining the estimate of free-energy barriers.
- Refined Free-Energy Landscape: This iterative process results in a highly refined and converged free-energy landscape, achieving accuracy that would be prohibitively expensive with REST2 or TREM alone.
Table 3: Key Software, Tools, and Methods for Convergence Analysis
| Tool / Resource | Type | Primary Function in Convergence Analysis |
|---|---|---|
| GROMACS [47] [1] | MD Software | High-performance MD engine with support for REST2 and TREM simulations. |
| AMBER, CHARMM, NAMD [1] [2] | MD Software | Alternative MD packages with implemented enhanced sampling methods. |
| CHARMM36 [47] | Force Field | A widely used and tested molecular force field for biomolecular simulations. |
| Markov Modeling (MSM) [47] [2] | Analysis Method | Infers thermodynamic and kinetic properties from many short simulations to assess convergence. |
| Denoising Diffusion Probabilistic Models (DDPM) [3] | AI/ML Model | A generative model that refines free-energy landscapes from replica exchange data. |
| Collective Variable (CV) | Analysis Concept | A low-dimensional descriptor of a slow conformational change, used for analysis and biasing. |
When applying these tools, researchers must be aware of common pitfalls. The quality of the force field is paramount, as inaccuracies will lead to sampling of incorrect conformations regardless of the sampling method's efficiency. Furthermore, convergence must be actively assessed, not assumed. Techniques include monitoring the stability of property estimates over time and checking for overlap between independent simulations [2]. For methods like REST2, careful parameterization of the replica ladder and scaled energy terms is critical to achieve high exchange rates and efficient sampling [1].
Sampling the conformational dynamics of biomolecules is fundamental to understanding their function, yet capturing rare transitions and converging ensembles remains a significant challenge in computational molecular biology. Traditional Temperature Replica Exchange Molecular Dynamics (T-REMD) has been a workhorse method but suffers from poor scaling with system size. Replica Exchange with Solute Tempering 2 (REST2) emerged as a Hamiltonian-based alternative that dramatically reduces computational requirements. More recently, Artificial Intelligence (AI)-based ensemble methods have introduced a paradigm shift by leveraging deep learning to sample conformational spaces without explicit physical simulation. This guide provides an objective comparison of these methodologies, examining their theoretical foundations, performance characteristics, and practical applications to inform researchers in structural biology and drug development.
Methodological Foundations
Temperature Replica Exchange MD (T-REMD)
T-REMD enhances sampling by running multiple replicas of the system in parallel at different temperatures. Replicas periodically attempt to exchange configurations based on a Metropolis criterion, allowing conformations to perform a random walk in temperature space and effectively overcome energy barriers. The number of replicas required scales with the square root of the system's degrees of freedom (f), making it computationally demanding for large, explicitly solvated systems [16].
Replica Exchange with Solute Tempering 2 (REST2)
REST2 addresses T-REMD's scaling issues by applying Hamiltonian scaling rather than temperature scaling. All replicas run at the same physical temperature, but the potential energy function for a selected "solute" region is scaled differently across replicas. The Hamiltonian for replica m is defined as:
E_m^REST2(X) = (β_m/β_0)E_pp(X) + √(β_m/β_0)E_pw(X) + E_ww(X)
where E_pp, E_pw, and E_ww represent solute-solute, solute-solvent, and solvent-solvent interaction energies respectively, β_m = 1/k_BT_m, and T_0 is the target temperature [16]. This approach focuses sampling effort on the region of interest, significantly reducing the number of replicas required compared to T-REMD.
AI-Based Ensemble Methods
AI-based approaches use deep learning models trained on structural databases and/or MD trajectories to directly generate conformational ensembles. These include:
- Generative models: Diffusion models and variational autoencoders learn to sample physically realistic conformations from simple noise distributions [3] [48].
- Coarse-grained ML potentials: Neural networks parameterize potential energy surfaces with reduced degrees of freedom [48].
- Hybrid AI-MD methods: AI models enhance MD sampling by identifying collective variables or generating starting structures [3].
These methods eschew traditional physical simulation in favor of data-driven pattern recognition and generation.
Performance Comparison
Computational Efficiency and Scaling
Table 1: Computational Efficiency and Resource Requirements
| Method | Replica/Resource Scaling | Typical Replica Count | Computational Advantage |
|---|---|---|---|
| T-REMD | Scales with √f (f = total degrees of freedom) [16] | 100+ for 72,000 atom system [9] | Simple implementation; well-established |
| REST2 | Scales with √fp (fp = solute degrees of freedom) [16] | ~16 for 72,000 atom system [9] | 3-10x fewer replicas vs. T-REMD [19] |
| AI Methods | Fixed cost per sample; no replicas needed [48] | No replicas | Statistically independent samples; GPU-optimized |
Sampling Performance and Applications
Table 2: Sampling Performance Across Biomolecular Systems
| Method | Small Protein Folding | IDP Conformational Ensembles | Binding Site Flexibility | Rare Event Sampling |
|---|---|---|---|---|
| T-REMD | Reliable for reversible folding [16] | Limited by system size [32] | Excellent with full flexibility [49] | Moderate (limited by temperature range) |
| REST2 | Excellent for β-hairpin and trpcage [16] | Efficient but may overcompact [9] | Targeted sampling of binding sites [19] | Good for local transitions |
| AI Methods | Varies by training data [48] | High diversity generation [32] | Limited atomic detail [48] | Limited to trained distributions |
Quantitative Performance Data
Table 3: Experimental Performance Metrics from Literature
| Study System | Method | Performance Metric | Result |
|---|---|---|---|
| Trpcage & β-hairpin | T-REMD | Reference folding sampling [16] | Baseline |
| Trpcage & β-hairpin | REST2 | Sampling efficiency vs T-REMD [16] | Greatly increased |
| p53 N-terminal domain | REST2 | Replica segregation issue [9] | Severe compaction |
| p53 N-terminal domain | REST3 | Improved temperature random walk [9] | Much more efficient |
| CLN025 mini-protein | REST2+DDPM | Accuracy vs T-REMD [3] | Comparable with fewer replicas |
| 82 test proteins | AlphaFlow | RMSF profile recovery [48] | Systematic improvement |
Experimental Protocols
Standard REST2 Implementation Protocol
The following protocol outlines a typical REST2 simulation setup in NAMD [19]:
System Preparation:
- Solvate the protein of interest in explicit water
- Select "hot region" atoms (typically the entire protein or binding site)
- Generate initial coordinates and PSF/PRMTOP files
Replica Parameterization:
- Determine temperature range (e.g., 300-500K)
- Calculate replica temperatures using:
T_i = T_0 * exp[ln(T_max/T_0)*(i/(N_rep-1))] - Typically 8-16 replicas for small to medium proteins [19]
Force Field Scaling:
- Scale bonded terms (dihedrals only, not bonds/angles)
- Scale Lennard-Jones ε parameters by (βm/β0)
- Scale atomic charges by √(βm/β0)
Simulation Parameters:
- Exchange attempt frequency: Every 1-2ps [19]
- Integration time step: 2fs
- Production run length: 100ns-1μs per replica
Analysis:
- Monitor acceptance probabilities (target: 20-40%)
- Check temperature random walks
- Compute conformational properties from lowest temperature replica
AI-Enhanced Sampling Protocol
A recently developed hybrid approach combines REST2 with diffusion models [3]:
Initial REST2 Simulation:
- Perform conventional REST2 simulation as above
- Collect configurations from all replicas
Diffusion Model Training:
- Train Denoising Diffusion Probabilistic Model (DDPM) on REST2 trajectories
- Learn joint probability distribution in configuration and rescaled potential energy space
Generative Sampling:
- Use trained DDPM to generate new conformations
- Apply importance sampling along known collective variables
Iterative Refinement:
- Use generated conformations to seed new REST2 simulations
- Focus sampling on high-barrier regions
- Repeat until convergence of free energy estimates
Research Reagent Solutions
Table 4: Essential Software Tools for Enhanced Sampling Studies
| Tool Name | Type | Primary Function | Key Features |
|---|---|---|---|
| NAMD | MD Engine | REST2 implementation [19] | Charm++ parallelism; Tcl scripting interface |
| GROMACS | MD Engine | T-REMD simulations [49] | GPU acceleration; REST2 via parameter modification |
| AlphaFlow | AI Ensemble Model | Conformational sampling [48] | Diffusion model; transferable across sequences |
| DiG (Distributional Graphormer) | AI Ensemble Model | Conformational sampling [48] | Graph neural network; recovers MD-observed states |
| VMD | Visualization | Analysis and visualization [19] | Hot region selection; trajectory analysis |
| AutoDock Vina | Docking | Ensemble docking [49] | Flexible ligand docking to multiple conformations |
Method Workflows
Limitations and Recent Advances
Known Limitations
REST2 Limitations:
- Promotes artificial protein conformational collapse at high effective temperatures [9]
- Can lead to replica segregation in temperature space for large IDPs [9]
- Sensitive to solute-solvent interaction scaling parameters [9]
T-REMD Limitations:
- Prohibitive replica count for large explicit solvent systems [16]
- Inefficient heating of solvent degrees of freedom irrelevant to protein conformational changes [16]
AI Method Limitations:
- Dependence on quality and diversity of training data [48]
- Limited ability to discover genuinely new conformational states [3]
- Challenges in ensuring thermodynamic faithfulness [48]
Emerging Solutions
REST3: A modified version addressing REST2's compaction issue by recalibrating solute-solvent van der Waals interactions to maintain appropriate chain expansion at high temperatures [9].
Hybrid AI-MD: Combining generative models with physical simulations to leverage strengths of both approaches [3]. For example, diffusion models can refine undersampled regions from REST2 simulations.
Transferable Coarse-Grained ML Potentials: Machine learning potentials trained on diverse protein simulations that maintain accuracy while accelerating sampling [48].
The comparative landscape of REST2, T-REMD, and AI-based ensemble methods reveals a complementary set of tools for biomolecular conformational sampling. T-REMD remains a reliable benchmark method but suffers from poor scaling. REST2 provides significant computational advantages for targeted sampling of specific regions, though it may require parameter optimization to avoid artifacts like artificial compaction. AI methods offer a fundamentally different approach with fixed computational cost and no replica management, but depend heavily on training data quality. The most promising future direction appears to be hybrid approaches that combine the thermodynamic rigor of physics-based methods like REST2 with the pattern recognition capabilities of AI models, leveraging the strengths of both paradigms to address the challenging problem of conformational sampling in complex biological systems.
Validating Ensembles Against Experimental Data for IDPs and Folded Proteins
Molecular dynamics (MD) simulations provide atomically detailed insights into the conformational ensembles of biomolecules, a capability crucial for understanding the function of both intrinsically disordered proteins (IDPs) and folded proteins [50] [51]. However, the accuracy of these simulations is highly dependent on the sampling method and the physical model, or force field, used [52]. Validating the resulting ensembles against experimental data is therefore an essential step to ensure their reliability. This guide compares the performance of Replica Exchange with Solute Tempering 2 (REST2), a widely used enhanced sampling method, against standard temperature replica exchange (TREM) and other alternatives, providing researchers with objective data to inform their methodological choices.
Methodological Foundations and Comparison
Standard Temperature Replica Exchange (TREM)
Principle: TREM overcomes quasi-ergodicity by running multiple parallel simulations (replicas) of the entire system at different temperatures. High-temperature replicas can cross energy barriers more easily, and periodic exchanges between replicas allow conformations sampled at high temperatures to propagate down to the temperature of interest [16] [45].
Limitations: The primary drawback is poor scalability with system size. The number of replicas required for efficient exchange scales as the square root of the system's degrees of freedom (√f). For proteins in explicit solvent, this makes TREM computationally prohibitive for large systems, as most replicas are expended on sampling the solvent rather than the solute [16] [5].
Replica Exchange with Solute Tempering 2 (REST2)
Principle: REST2 is a Hamiltonian replica exchange method that focuses the enhanced sampling on the solute. Instead of changing the temperature, REST2 scales the Hamiltonian for different replicas. All replicas run at the same physical temperature, but the potential energy terms associated with the solute are scaled down in higher replicas, effectively lowering energy barriers for the protein while the solvent remains "cold" [16] [45].
Key Differentiator: This approach bypasses TREM's poor system-size scaling. The number of required replicas scales with the square root of the solute's degrees of freedom (√fp), drastically reducing the number of parallel processes needed for solvated systems [16]. A key improvement of REST2 over its predecessor (REST1) is its Hamiltonian scaling, which enhances sampling efficiency for large-scale conformational changes [16].
Other Advanced Sampling Methods
- Hybrid Tempering (REHT): An extension of REST2 that also slightly heats the solvent bath. This "rewiring" of the hydration shell helps the solute cross large entropic barriers, further improving sampling efficiency for complex landscapes like those of metamorphic proteins and IDPs [5].
- HREST-BP: Combines REST2's global solute scaling with targeted biasing potentials along specific collective variables (CVs). This is particularly useful for enhancing sampling along known reaction coordinates, such as glycosidic linkages in carbohydrates [45].
The workflow below illustrates the core REST2 process and its key advantage in Hamiltonian scaling.
Quantitative Performance Comparison
The following tables summarize key performance metrics from published studies comparing REST2, TREM, and other methods across various protein systems.
Table 1: Sampling Efficiency for Protein Folding
| Protein System | Method | Replicas Used | Time to Fold (ns) | Folding Free Energy Barrier (kcal/mol) | Key Observation |
|---|---|---|---|---|---|
| Trpcage (20 residues) | TREM | 16-24 [16] | >300 [5] | ~2.1 (Ref. [5]) | Prohibitively high CPU cost [16] |
| REST2 | 8 [16] | ~100 [5] | ~2.0 [5] | Efficient sampling of folded/unfolded states [16] [5] | |
| β-Hairpin | TREM | 16-24 [16] | >300 [5] | N/R | Poor scalability with explicit solvent [16] |
| REST2 | 8 [16] | ~100 [5] | N/R | Greatly reduced CPU requirement [16] | |
| Performance Summary | REST2 Advantage | ~66% Fewer | ~3x Faster | Accurate | Enables ab initio folding in explicit water [16] |
Table 2: Application to Complex Biomolecular Systems
| System Type | Method | Key Performance Metric | Agreement with Experiment |
|---|---|---|---|
| N-glycans (HIV gp120) | HREST-BP (REST2 + CV bias) [45] | Efficient sampling of coupled local linkages and long-range motions with only 6-8 replicas [45]. | Recapitulated known conformational properties of complex saccharides [45]. |
| Histatin-5 (IDP) | REHT (REST2 hybrid) [5] | Accurate ensemble generation without the need for reweighting [5]. | NMR and SAXS data matched well with calculated ensemble averages [5]. |
| RFA-H (Metamorphic) | REHT (REST2 hybrid) [5] | Successful mapping of multi-funneled free energy landscape [5]. | Generated ensembles matched various biophysical experiments [5]. |
Experimental Validation Protocols
Simulated conformational ensembles must be validated against empirical data. The table below lists key experimental techniques used for this purpose.
Table 3: Research Reagent Solutions for Ensemble Validation
| Research Reagent / Technique | Function in Validation |
|---|---|
| Nuclear Magnetic Resonance (NMR) Spectroscopy | Provides atomistic data on local structure (chemical shifts), long-range contacts (NOEs), and dynamics (relaxation) for proteins in solution [33] [51]. |
| Small-Angle X-Ray Scattering (SAXS) | Yields low-resolution data on the global size and shape (radius of gyration, Rg) of the ensemble in solution [33] [51]. |
| Förster Resonance Energy Transfer (FRET) | Measures distance distributions between specific sites on a protein, reporting on global compaction and dynamics [53]. |
| Circular Dichroism (CD) Spectroscopy | Provides information on the secondary structure composition (e.g., helical, beta-sheet, random coil) of the ensemble [32]. |
The Validation Workflow: From Simulation to Experiment
A robust validation protocol involves using "forward models" to calculate experimental observables from the simulated ensemble and iteratively comparing them to the actual data [33] [51]. The diagram below illustrates this integrative workflow.
Maximum Entropy Reweighting
When initial simulations do not fully agree with experiments, the maximum entropy reweighting procedure provides a statistically sound method to refine the ensemble [33] [51]. This approach minimally adjusts the statistical weights of conformations in the original simulation to achieve agreement with experimental data while maximizing the entropy of the resulting ensemble—meaning it is the least biased adjustment possible [33]. A fully automated version of this method has been shown to produce accurate, force-field-independent conformational ensembles of IDPs when sufficient experimental data is available [33].
Protocol:
- Run extended unbiased or enhanced sampling MD simulations with a chosen method (e.g., REST2).
- Calculate experimental observables for every frame in the ensemble using validated forward models.
- Determine optimal weights for each frame by minimizing the discrepancy with experimental data under the maximum entropy constraint.
- Analyze the reweighted ensemble and compute the Kish ratio to ensure the ensemble is not overfit (i.e., that a sufficient number of conformations contribute significantly to the final ensemble) [33].
For simulating conformational ensembles of proteins in explicit solvent, REST2 and its derivatives offer a superior balance of computational efficiency and sampling power compared to standard TREM. The key advantage of REST2 is its focused sampling on the solute, which drastically reduces the required computational resources and makes the ab initio folding of proteins and the sampling of complex IDP landscapes feasible [16] [5]. Ultimately, the choice of method should be guided by the biological question. For studies requiring atomic detail in explicit solvent, REST2 is the recommended starting point. The resulting ensembles must be rigorously validated against a suite of experimental data, such as NMR and SAXS, with maximum entropy reweighting providing a powerful tool to reconcile simulations and experiments into a single, accurate conformational ensemble [33] [51].
Conclusion
The comparative analysis unequivocally demonstrates that REST2 represents a significant leap forward in conformational sampling, offering a more computationally efficient pathway than standard MD or T-REMD for explicit solvent simulations of biomolecules. By strategically scaling the Hamiltonian of a solute region, REST2 dramatically reduces the number of required replicas and accelerates the exploration of complex free energy landscapes, particularly for protein folding and IDP studies. However, practitioners must be aware of its nuances, such as the potential for artificial compaction in IDPs, which is being addressed by next-generation protocols like REST3. The future of biomolecular simulation lies in the intelligent integration of such enhanced sampling methods with AI-driven approaches and high-performance computing. For biomedical research, this convergence promises to unlock a deeper understanding of protein function, enable the targeting of previously 'undruggable' proteins with conformational flexibility, and ultimately accelerate the design of novel therapeutics.
References
- 1. Feasibility of Combining Biomolecular Conformational ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12308781/]
- 2. Computational methods for exploring protein conformations [https://pmc.ncbi.nlm.nih.gov/articles/PMC7458412/]
- 3. Hamiltonian replica exchange augmented with diffusion ... [https://arxiv.org/html/2505.08357v2]
- 4. Residence time in drug discovery: current insights and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12241192/]
- 5. High resolution ensemble description of metamorphic and ... [https://www.nature.com/articles/s41467-021-21105-7]
- 6. Molecular Dynamics - GROMACS 2025.4 documentation [https://manual.gromacs.org/documentation/current/reference-manual/algorithms/molecular-dynamics.html]
- 7. When Does Molecular Dynamics Improve RNA Models ... [https://www.sciencedirect.com/science/article/pii/S2001037025004040]
- 8. Advances in Molecular Dynamics Simulations and ... [https://www.mdpi.com/1422-0067/21/17/6339]
- 9. Re-balancing Replica Exchange with Solute Tempering for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10795075/]
- 10. Full structural ensembles of intrinsically disordered ... [https://www.nature.com/articles/s42003-021-01759-1]
- 11. RNA Dynamics and Interactions Revealed through Atomistic Simulations [https://arxiv.org/html/2507.08474v1]
- 12. Simulated Solute Tempering 2: An Efficient and Practical ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12661556/]
- 13. Molecular dynamics simulations illuminate the role of ... [https://elifesciences.org/reviewed-preprints/102410]
- 14. Enhanced sampling of protein conformational changes via ... [https://www.nature.com/articles/s41467-025-55983-y]
- 15. How good is generative diffusion model for enhanced ... [https://pubmed.ncbi.nlm.nih.gov/40960922/]
- 16. Replica Exchange with Solute Scaling: A more efficient ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3172817/]
- 17. Hamiltonian replica exchange augmented with diffusion-based generative models and importance sampling to assess biomolecular conformational basins and barriers [https://arxiv.org/html/2505.08357v1]
- 18. A Standardized Benchmark for Machine-Learned ... [https://arxiv.org/abs/2510.17187]
- 19. A generic implementation of replica exchange with solute ... [https://www.sciencedirect.com/science/article/abs/pii/S001046551500329X]
- 20. Effects of forcefield and sampling method in all-atom simulations of... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5638406/]
- 21. Structure and dynamics in drug discovery [https://www.nature.com/articles/s44386-024-00001-2]
- 22. NAMD 2.13 New Features [https://www.ks.uiuc.edu/Research/namd/2.13/features.html]
- 23. Extended Sampling of Macromolecular Conformations from Uniformly... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11672670/]
- 24. Statistical accuracy of molecular dynamics-based methods for... [https://pubs.rsc.org/en/content/articlelanding/2024/cp/d4cp02564d]
- 25. NAMD GPU Benchmarks and Hardware Recommendations [https://www.exxactcorp.com/blog/benchmarks/namd-gpu-benchmarks-and-hardware-recommendations]
- 26. NAMD [https://www.osc.edu/resources/available_software/software_list/namd]
- 27. Folding of Trp-cage Mini Protein Using Temperature and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2672022/]
- 28. (PDF) Crystal and NMR structures of a Trp - cage mini-protein... [https://www.academia.edu/83921712/Crystal_and_NMR_structures_of_a_Trp_cage_mini_protein_benchmark_for_computational_fold_prediction]
- 29. Hamiltonian Replica Exchange Augmented with Diffusion ... [https://pubmed.ncbi.nlm.nih.gov/41165096/]
- 30. Simulated Solute Tempering 2: An Efficient and Practical ... [https://pubmed.ncbi.nlm.nih.gov/41139839/]
- 31. simulation of Ab by replica exchange with... initio folding Trpcage [https://pubmed.ncbi.nlm.nih.gov/18775599/]
- 32. Use of AI-methods over MD simulations in the sampling ... [https://www.frontiersin.org/journals/molecular-biosciences/articles/10.3389/fmolb.2025.1542267/full]
- 33. Determining accurate conformational ensembles of ... [https://www.nature.com/articles/s41467-025-64098-3]
- 34. The impact of molecular dynamics on drug design ... [https://www.sciencedirect.com/science/article/abs/pii/S1359644615000215]
- 35. Sampling conformational space of intrinsically disordered ... [https://www.sciencedirect.com/science/article/abs/pii/S109332631630496X]
- 36. Re-Balancing Replica Exchange with Solute Tempering for ... [https://pubmed.ncbi.nlm.nih.gov/36791464/]
- 37. Temperature-induced collapse of a disordered peptide ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5966312/]
- 38. Artificial intelligence guided conformational mining of ... [https://www.nature.com/articles/s42003-022-03562-y]
- 39. Error and efficiency of replica exchange molecular dynamics ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2780465/]
- 40. ACES: Optimized Alchemically Enhanced Sampling - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC10333454/]
- 41. Inhibitory mechanisms of amentoflavone on amyloid-β ... [https://www.nature.com/articles/s41598-025-10623-9]
- 42. Molecular Simulations Integrated with Experiments for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11242915/]
- 43. Improved Prediction of Molecular Response to Pulling by ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10823473/]
- 44. Fouling behavior of protein in mixed solvent systems [https://www.sciencedirect.com/science/article/pii/S0376738823010177]
- 45. Enhanced Conformational Sampling Using Replica Exchange ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4463548/]
- 46. Replica exchange with solute scaling: a more efficient version ... [https://pubmed.ncbi.nlm.nih.gov/21714551/]
- 47. Deciphering allosteric mechanisms in KRAS activation: insights from... [https://pubs.rsc.org/en/content/articlehtml/2025/ra/d4ra07924h]
- 48. AI-based Methods for Simulating, Sampling, and Predicting ... [https://arxiv.org/html/2509.17224v1]
- 49. Supercomputer- Based Docking Drug Discovery Pipeline... Ensemble [https://pmc.ncbi.nlm.nih.gov/articles/PMC7754786/]
- 50. Atomistic molecular dynamics simulations of intrinsically ... [https://www.sciencedirect.com/science/article/pii/S0959440X25000478]
- 51. Using simulation to interpret experimental data in terms of ... [https://www.sciencedirect.com/science/article/abs/pii/S0959440X16302056]
- 52. Validating Molecular Dynamics Simulations Against ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6420231/]
- 53. Towards Accurate Coarse-Grained Simulations of Disordered ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9910785/]